Find closest 8-connected chessboard distance between multiple pairs of points: shortest m-path
An answer to this question on Stack Overflow.
Question
I am working on binary images in Python using OpenCV. I have two sets of points: PNodes and FNodes. I want to find the closest PNode to each of the FNodes (shortest m-path); closest in terms of 8-connected chessboard distance.
In the example below, suppose PNodes (donated by *) are: (6,1), (6,5) and (5,8). (indexing starts from 0, first element is row number). FNodes (denoted by #) are: (0,1), (0,9), (1,6), (2,5) and (4,3).
import numpy as np
In = np.array ((
[ 0, 1#, 0, 0, 0, 0, 0, 0, 0, 1#, 0],
[ 1, 0, 0, 0, 0, 0, 1#, 0, 1, 0, 0],
[ 0, 1, 0, 0, 0, 1#, 0, 0, 1, 0, 0],
[ 0, 0, 1, 0, 0, 0, 1, 0, 0, 1, 0],
[ 0, 1, 1, 1#, 0, 1, 0, 0, 1, 0, 0],
[ 0, 1, 0, 0, 0, 1, 0, 0, 1*, 0, 0],
[ 0, 1*, 0, 0, 0, 1*, 0, 0, 1, 0, 0],
[ 0, 1, 0, 0, 1, 0, 1, 1, 0, 0, 0],
[ 0, 0, 1, 1, 0, 0, 0, 1, 0, 0, 0]), dtype = "uint8")
Distance_Matrix = np.array ((
[ 0, 6#, 0, 0, 0, 0, 0, 0, 0, 5#, 0],
[ 5, 0, 0, 0, 0, 0, 5#, 0, 4, 0, 0],
[ 0, 4, 0, 0, 0, 4#, 0, 0, 3, 0, 0],
[ 0, 0, 3, 0, 0, 0, 3, 0, 0, 2, 0],
[ 0, 2, 2, 3#, 0, 2, 0, 0, 1, 0, 0],
[ 0, 1, 0, 0, 0, 1, 0, 0, **, 0, 0],
[ 0, **, 0, 0, 0, **, 0, 0, 1, 0, 0],
[ 0, 1, 0, 0, 1, 0, 1, 1, 0, 0, 0],
[ 0, 0, 1, 1, 0, 0, 0, 1, 0, 0, 0]), dtype = "uint8")
I am not concerned with the exact value of distance, I just want to find out the closest pair. Something like this: FNode at (0,1) is closest to PNode at (6,1). FNode at (4,3) is closest to PNode at (6,1). All distances are in terms of 8-connected chessboard distance.
Ultimate requirement from this entire process: Basically, I just want to make sure all PNodes have atleast 1 FNode which lie within a given distance range (along the path of 1s).
Suppose PNode (PN_1) has a FNode (FN_1) which lies within the required distance range, I also make sure that PN_1 is closest to FN_1, and not any other PNode.
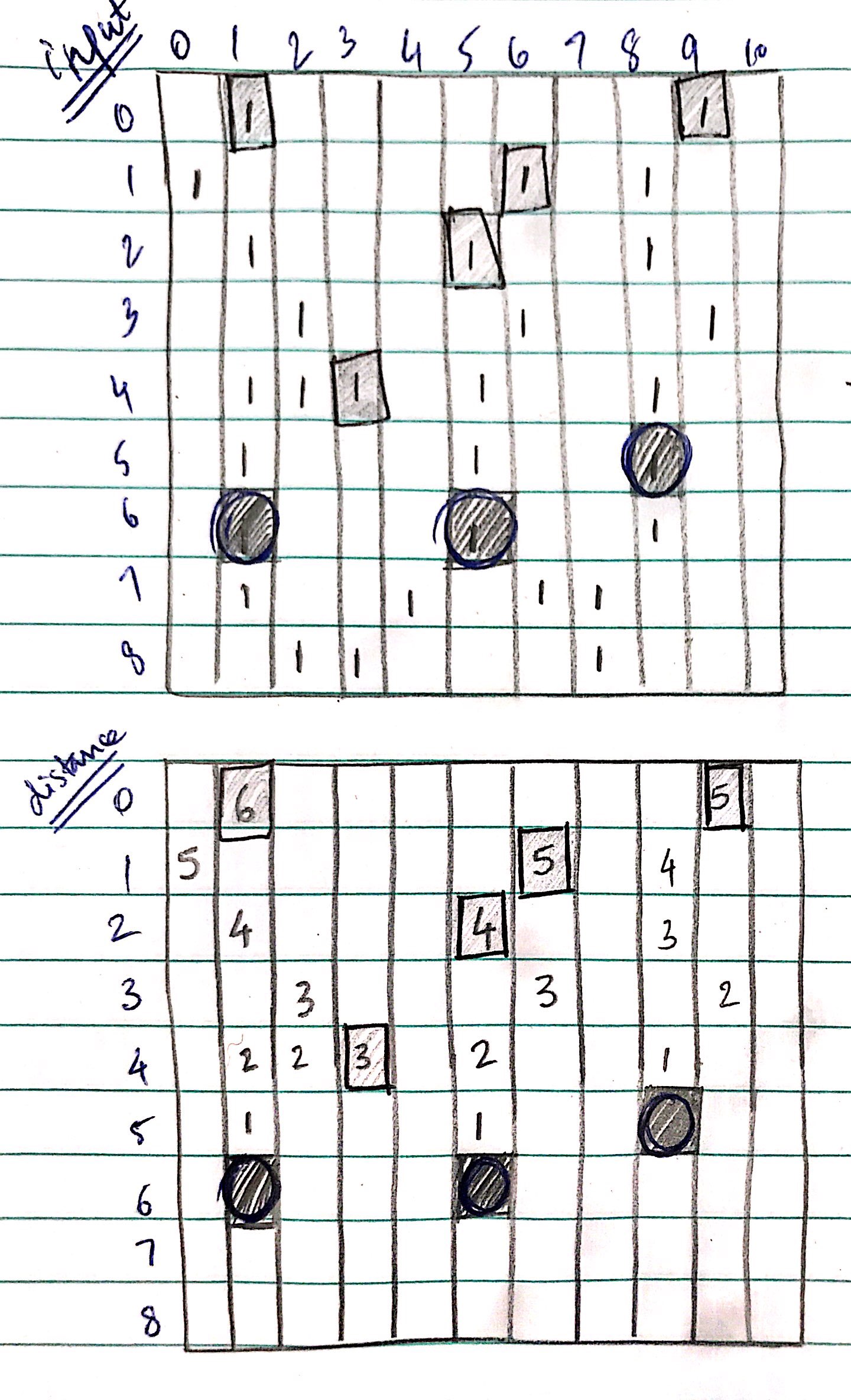
For a better understanding, I have attached an image below; FNodes are rectangular and PNodes are circular.
I don't care about other elements in this matrix, apart from those of PNodes and FNodes, as depicted.
Answer
I'd use Dijkstra's algorithm for this. It finds the shortest distance between a source node and every other node by exploring nearer nodes before farther nodes.
Seed a Dijkstra search at each of your P-nodes and halt the search when you either find the first F-node or exceed your distance constraint.
Since you don't have a graph, per se, you can generate neighbours using offsets. For instance if you have a cell (x,y) you can define some offsets
dx = [-1,-1, 0, 1,1,1,0,-1]
dy = [ 0,-1,-1,-1,0,1,1, 1]
and then take (x+dx[i], y+dy[i]) for i∈[0,7] to generate the neighbours of a given cell.
You can define a separate 2D array/matrix for tracking whether cells have been visited.