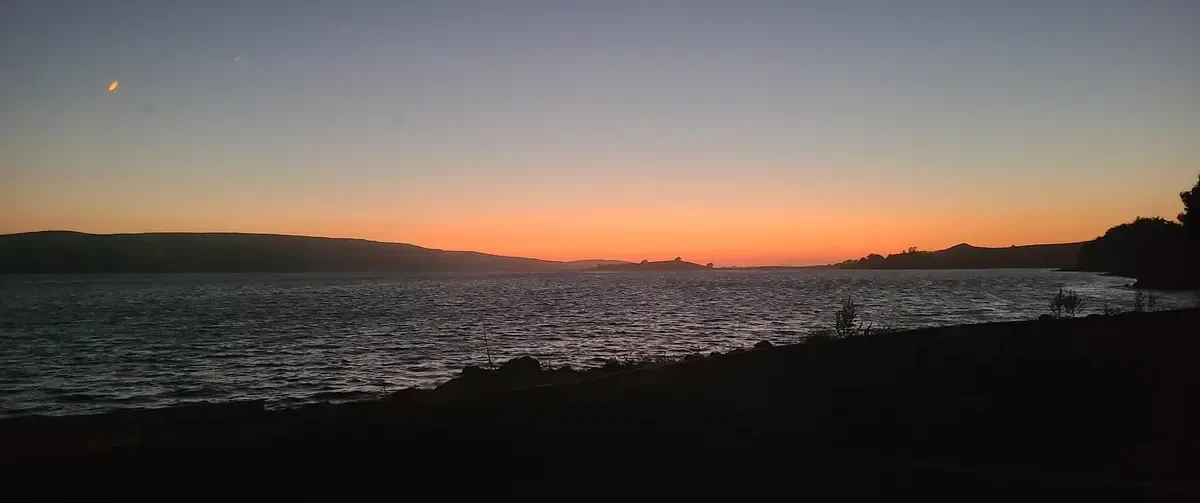
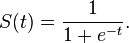Tomales Bay is one of the best places in California to kayak through bioluminescent plankton, but you can only see it if three things align: it has to be dark enough (moon below the horizon), late enough (after full darkness), and the right season (late spring through fall when dinoflagellate populations peak). What follows is a trip report followed by some planning tools for your own trip.

The First Trip — September 23–24, 2022
We reserved Boat Site B at Point Reyes National Seashore Campground, a group site on the west shore of Tomales Bay at Marshall Beach, accessible only by water.
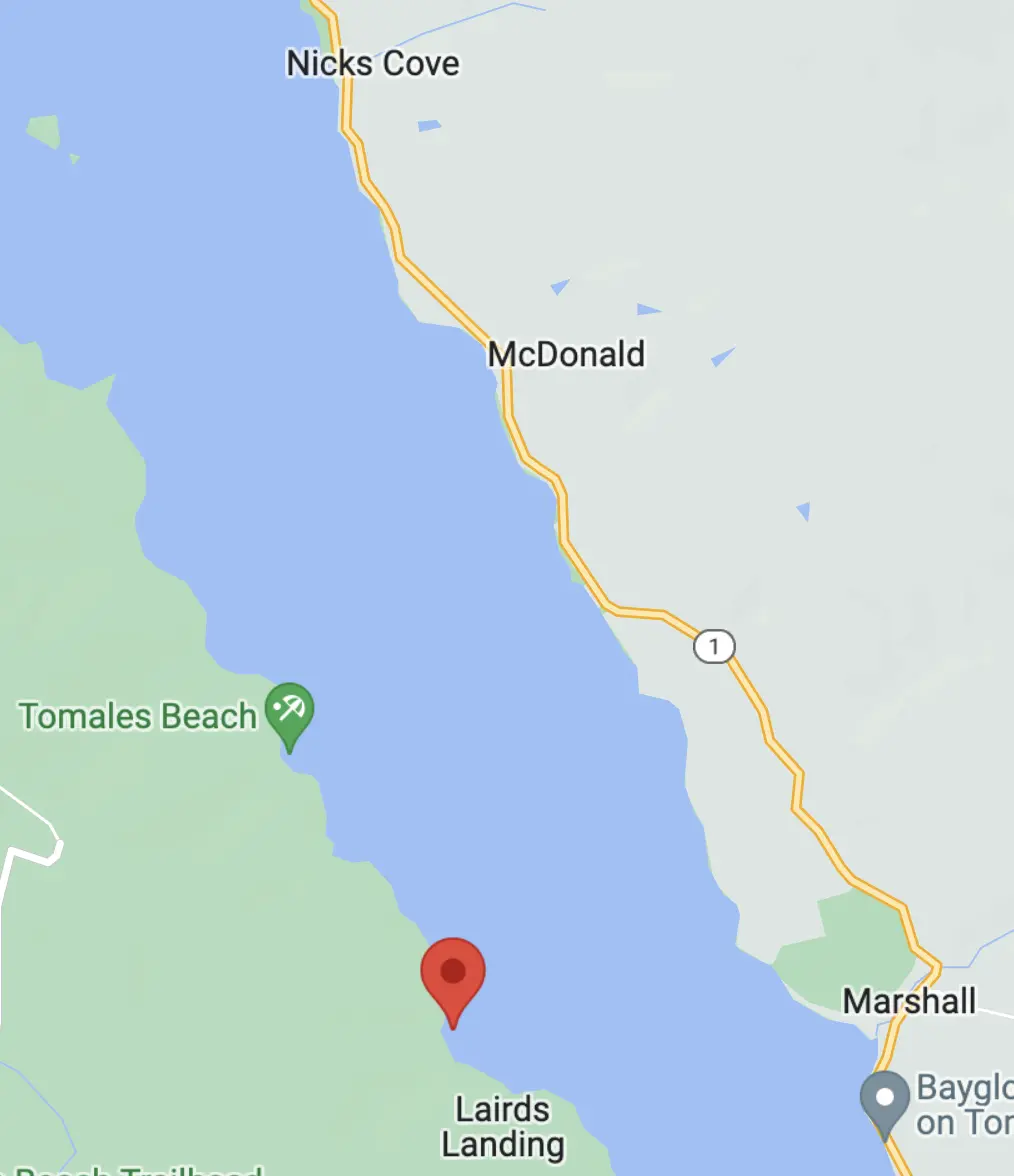
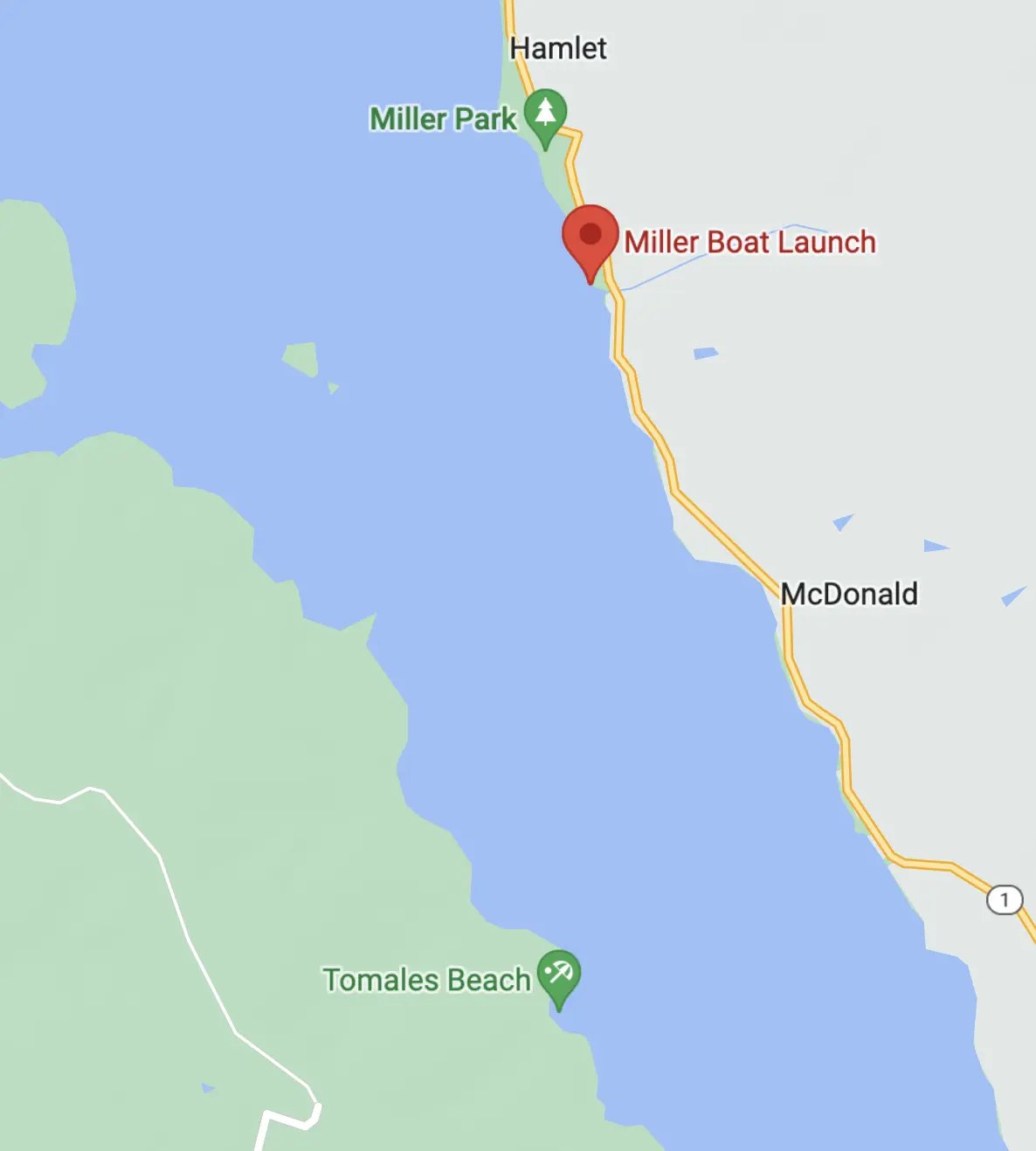
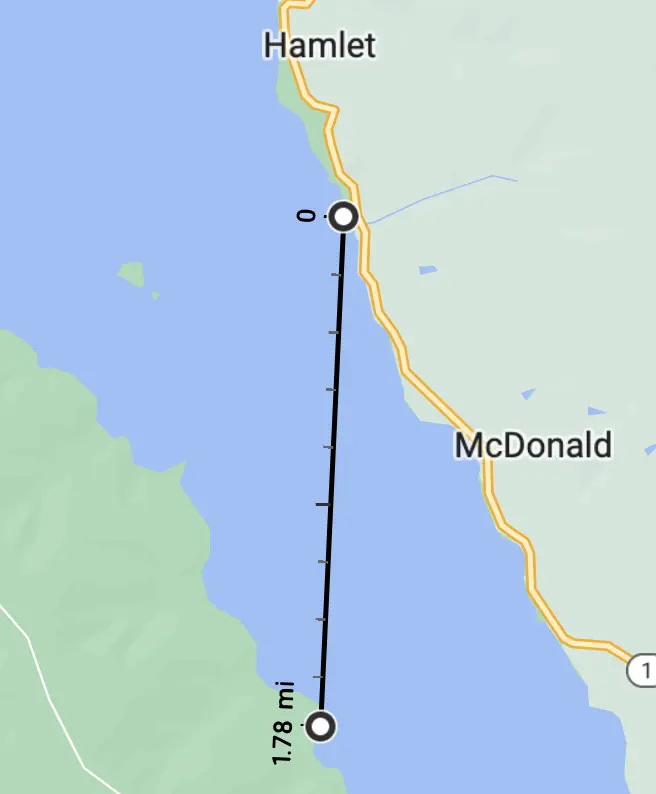
The original plan was to park overnight at, and launch from, Miller Boat Launch on the east shore and paddle across to camp.
Straight-line, the route is about 1.78 miles.
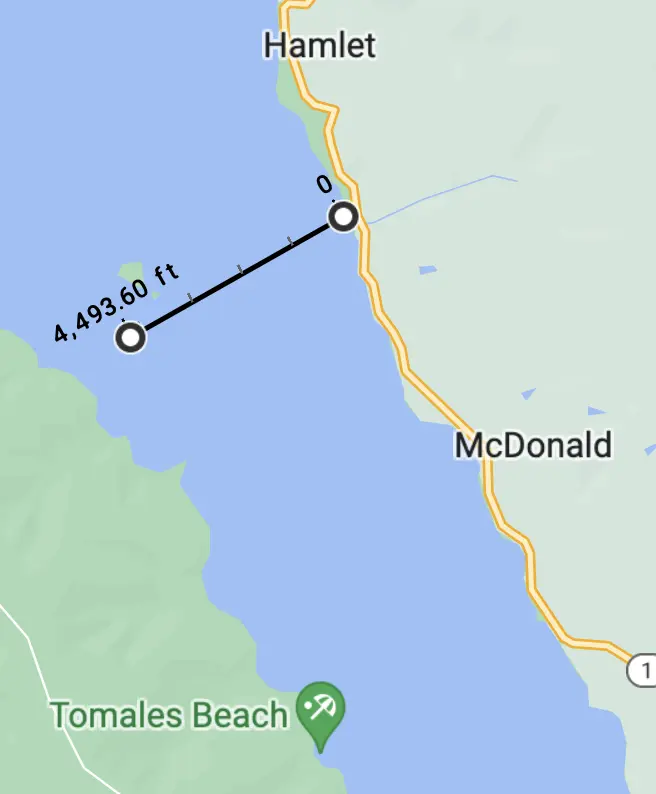
By timing the crossing for slack tide, though, we could have cut straight across the Bay — only about 4,500 feet — and then head southeast along the shore to the camp site. This would have minimized exposure to open water risks.
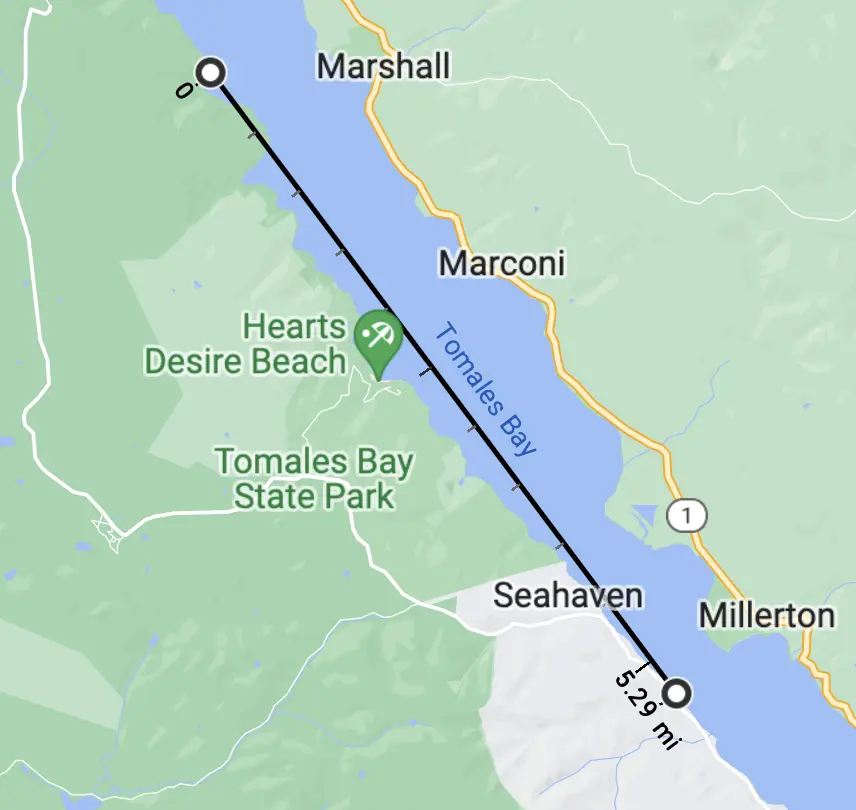
But, in planning this, I chickened out a bit. The group included people with variable kayaking experience and high afternoon winds can lead to large waves on the Bay, so we instead launched from Chicken Ranch Beach to the south, and paddled 5.29 miles along the shore so that there'd be an easy way to bail if things went wrong. This choice brought us into Type II Fun territory.
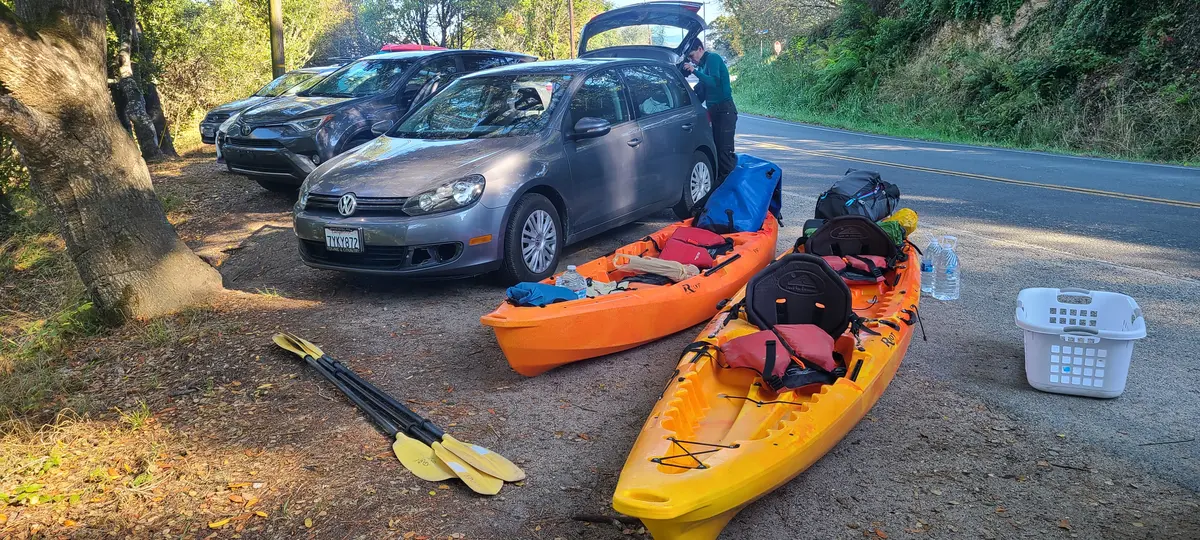
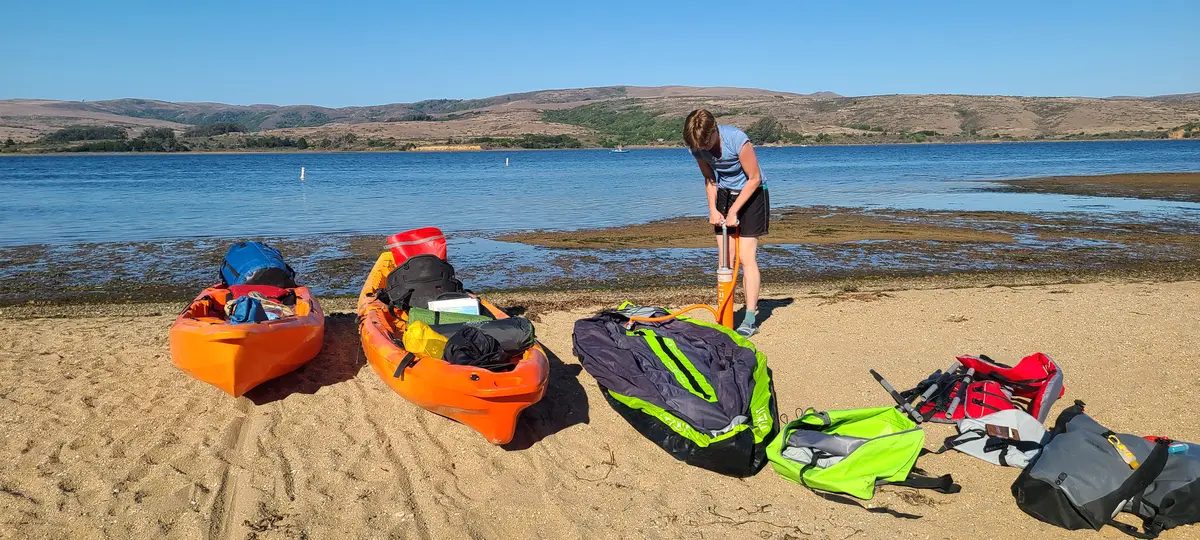



I chose the date to maximize darkness. Moonset on September 23 was at 6:19 PM — well before sunset — leaving the whole evening dark. (The planner below includes all this information in an easy-to-use form.)
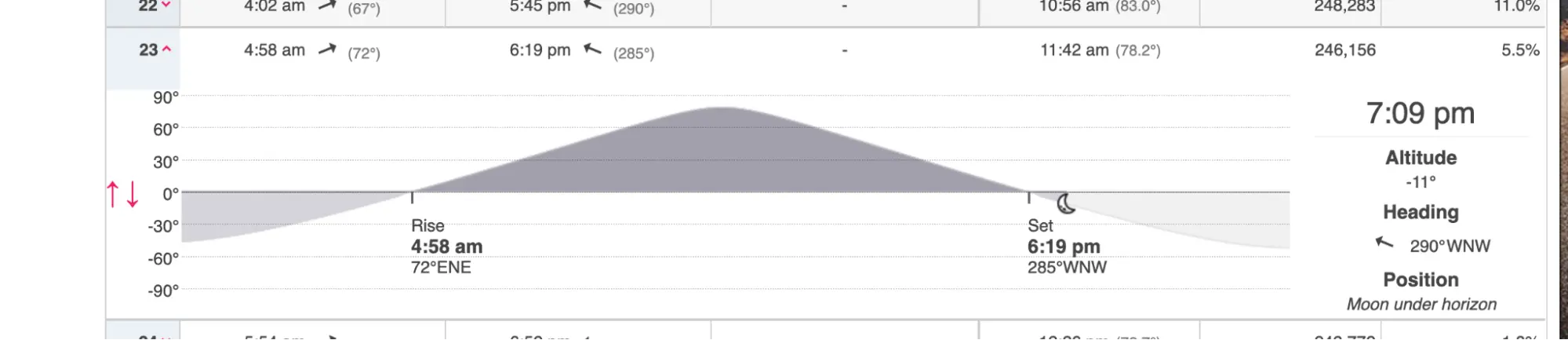
In addition, civil twilight ended at 7:39 PM, nautical twilight at 8:06 PM, and astronomical twilight at 8:33 PM.
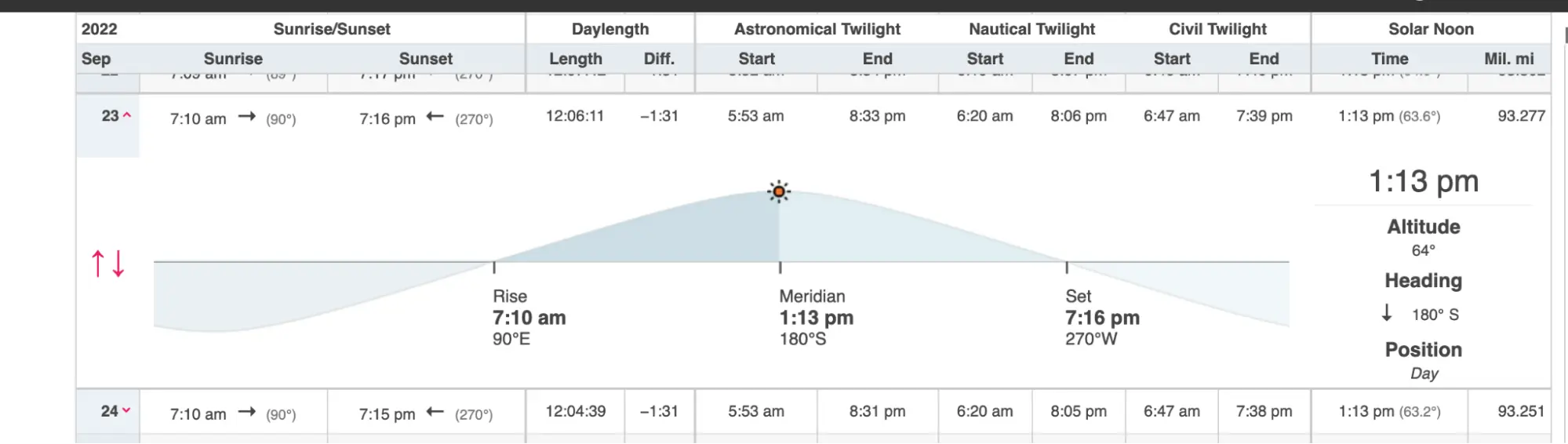
Winds pick up in the afternoons so launching before noon is recommended and, in fact, local outfitters won't rent kayaks after noon even though winds tend to drop off again later in the day. Given the tide and light, starting around 4:00 PM during slack tide should have given enough time to paddle and set up camp before dark.
However, due to folks' work schedules, we weren't able to get onto the water until 5:12 PM (a vanguard from our group acquired the kayaks). This meant meant fighting the incoming tide that started at 5:53 PM on the 23rd. But not just the tide! Also that wind. The result was a brutal, soaking paddle. We got to camp at 7:17 PM, so the trip took about two hours.
On the way out we didn't time it much better and got on the water at 11:25 AM to fight the outgoing tide on our way back. The winds were calm and the paddling was much easier than the day before. We took it slow and arrived at 1:18 PM, so the trip took about two hours.
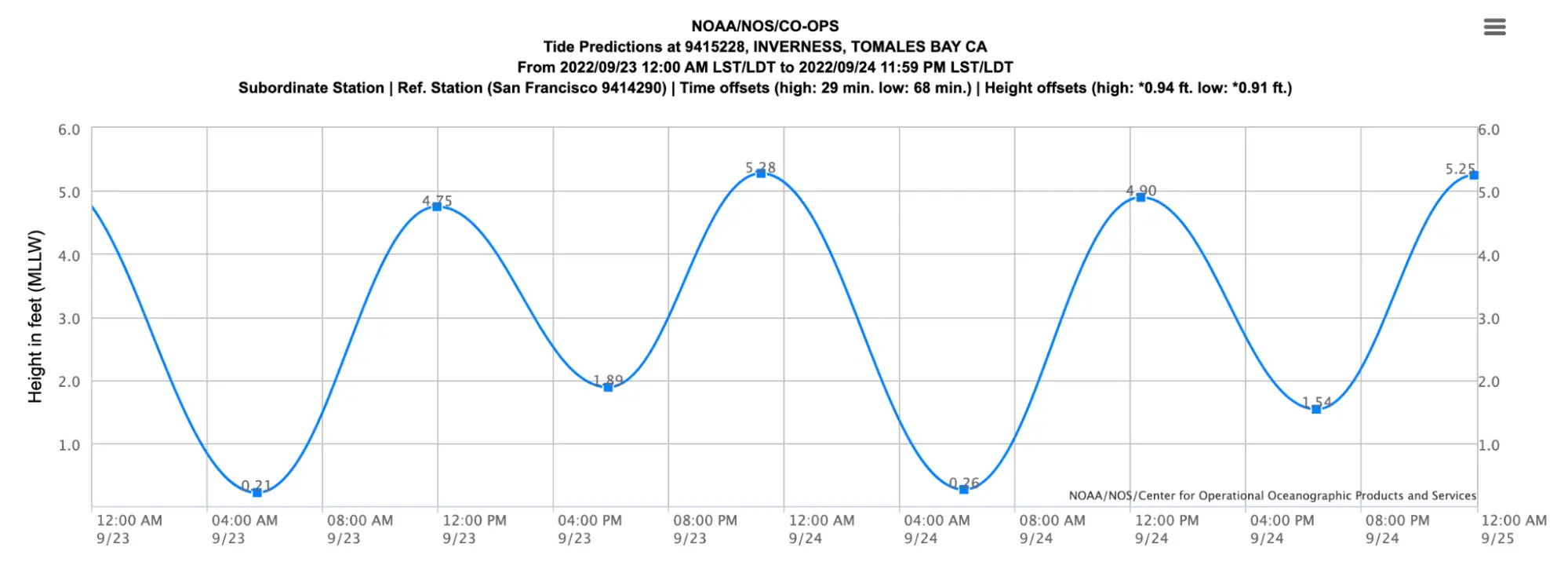
Most beaches on the west shore are tidally dependent and will disappear at tides above 5 ft. Since September 23 had a high tide of 5.28 ft at 11:11 PM, we needed to set up camp well above the waterline.

Cold and wet, some members of the trip retreated to tents immediately to get warm and no one was up for venturing back out again into the dark. The fabled bioluminescence didn't show itself in the bay near our tents.
But the weekend wasn't a wash. Our visit corresponded with the annual campout of the Traditional Small Craft Association. They played folk music by their fire well into the night.



The next morning we went hiking up and around our mini-bay.
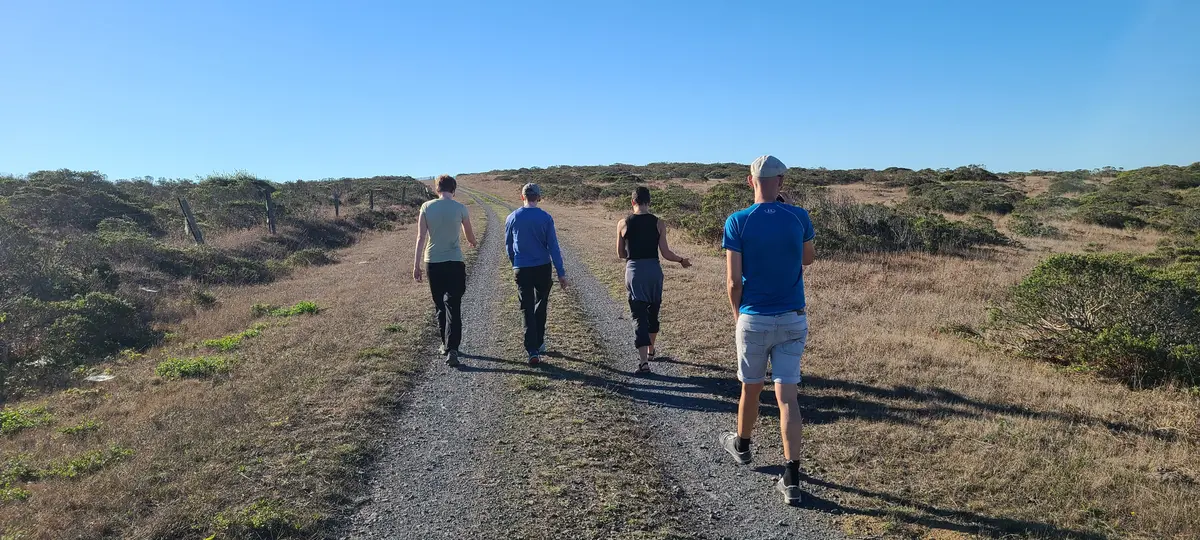
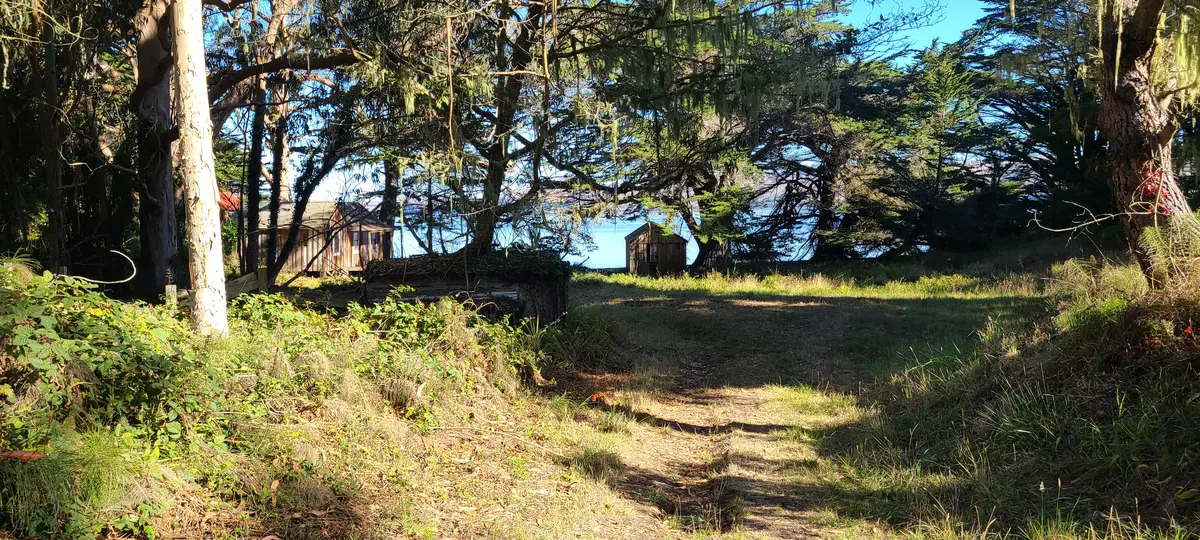


It took a while, so we investigated climbing across the cliffs to get back to camp.
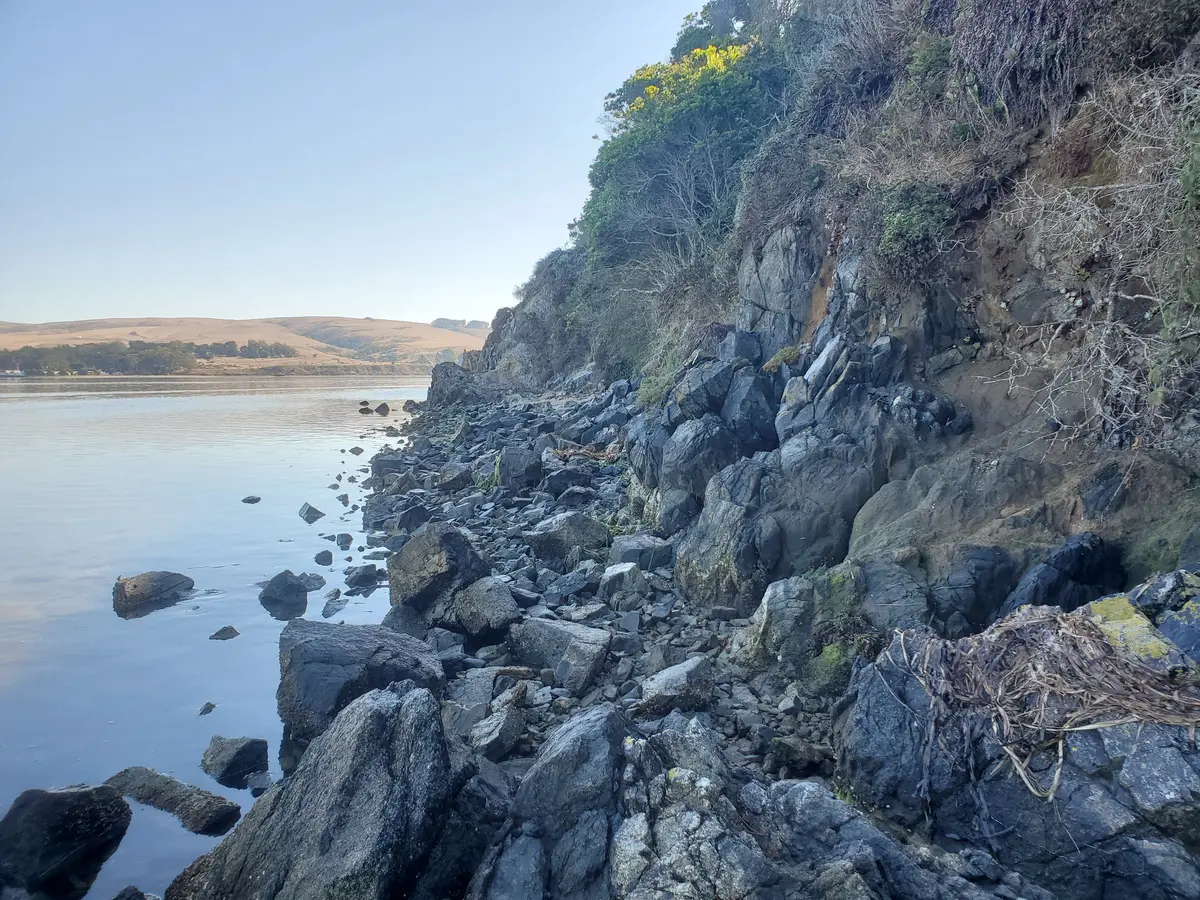

But were spared that adventure by the TCSA coming over in one of their boats!
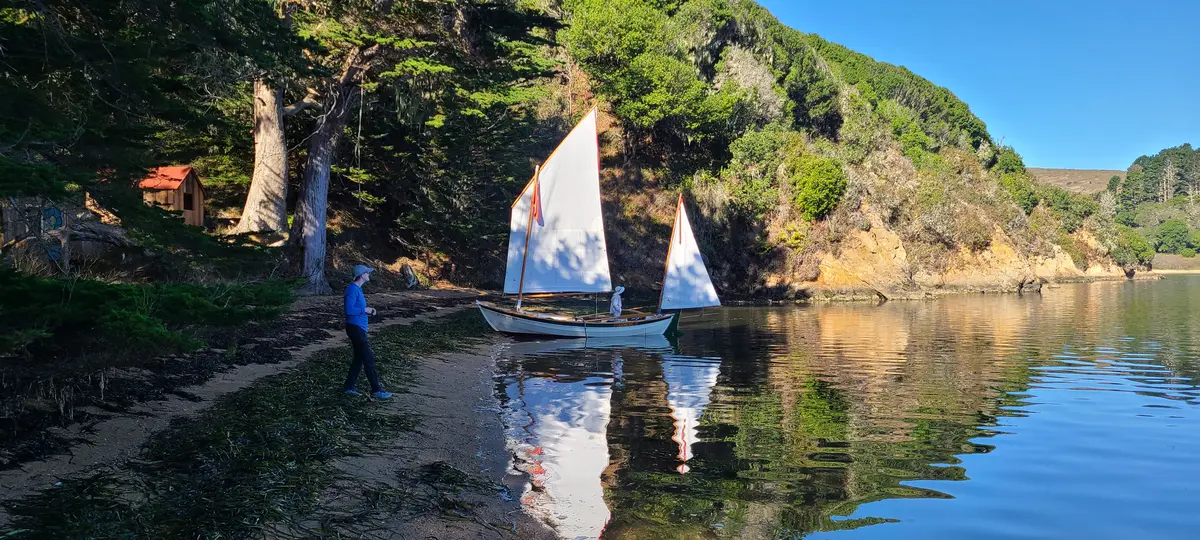
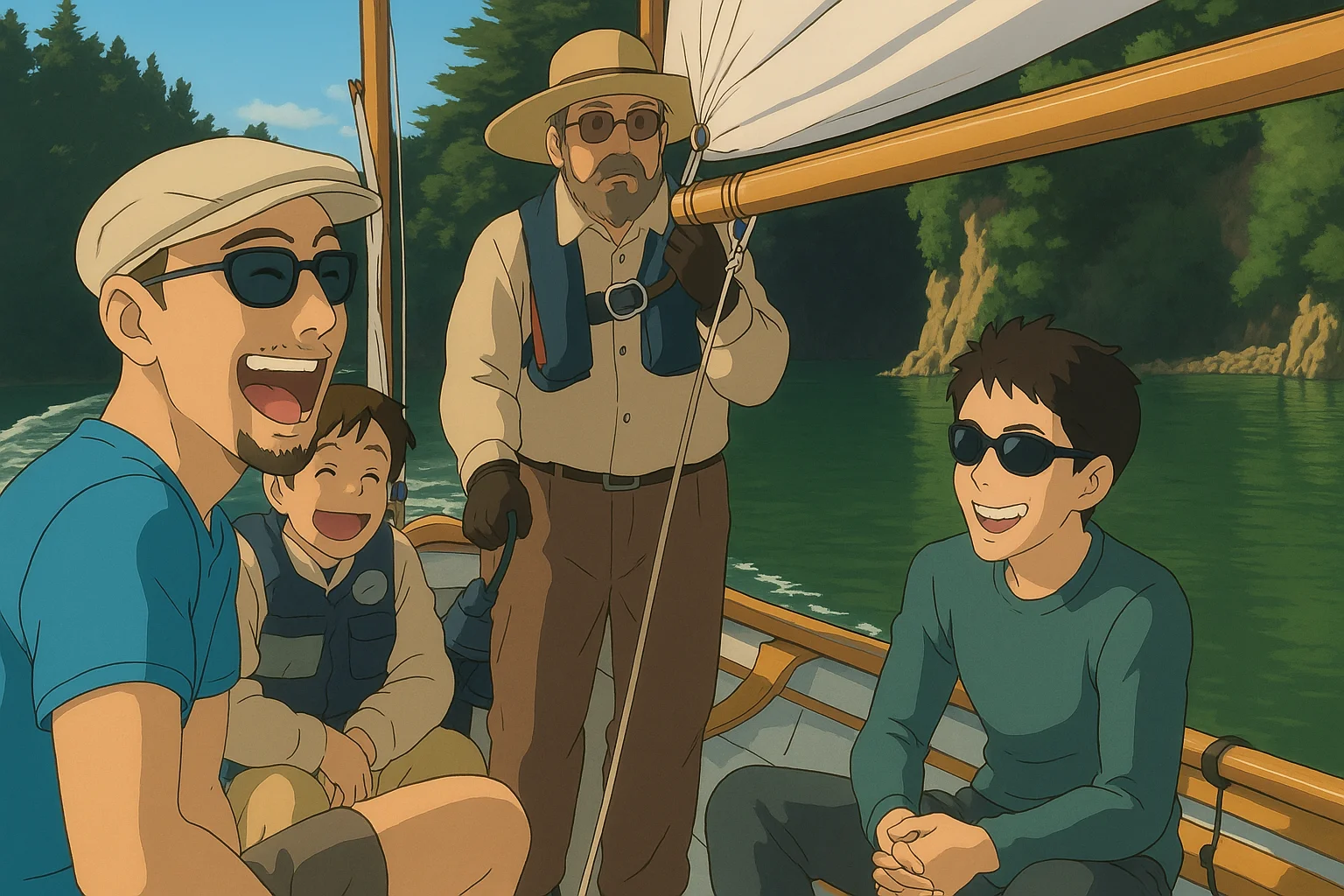
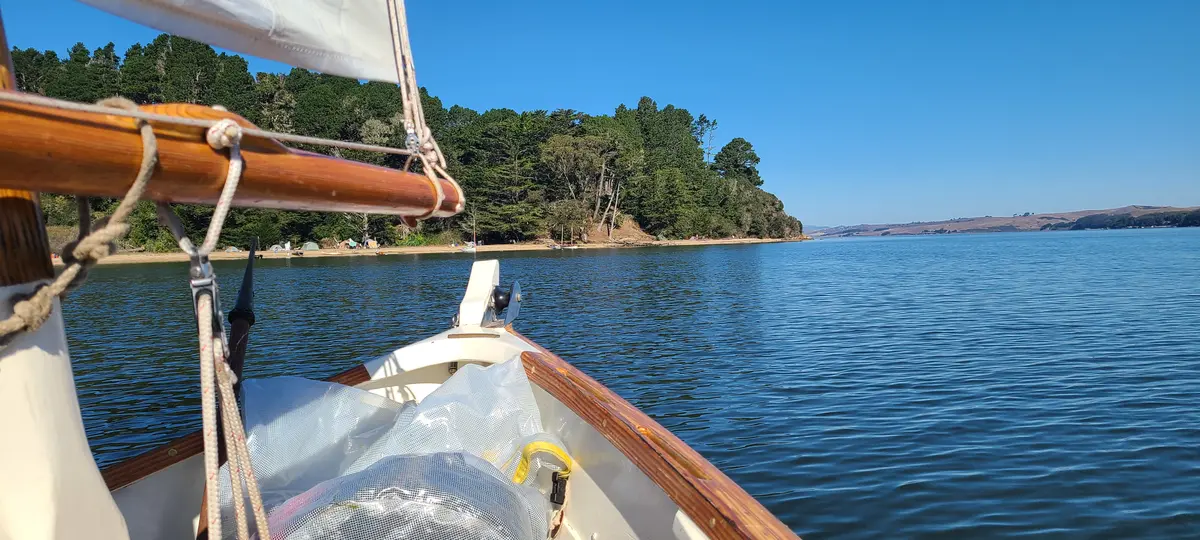
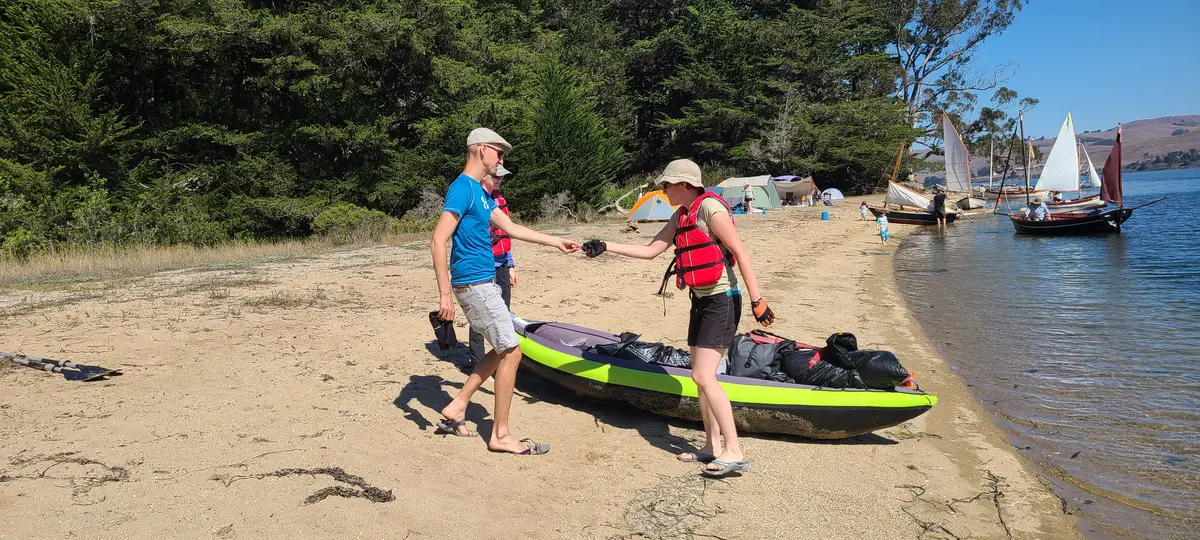




The Second Trip — October 6–7, 2023
I returned a year later with a friend to try again. Blue Waters normally rents sit-on-top kayaks, but as a former kayak guide I was able to get a lighter enclosed double. This time we used the Miller Boat Launch. The winds were, by luck, exceptionally calm when we got on the water at 3:30 PM. By 4:39 PM we'd successfully crossed to the opposite shore and were making our way along it and by 5:47 PM we had our tent set up at Tomales Beach (which is closer to Miller than Marshall Beach).
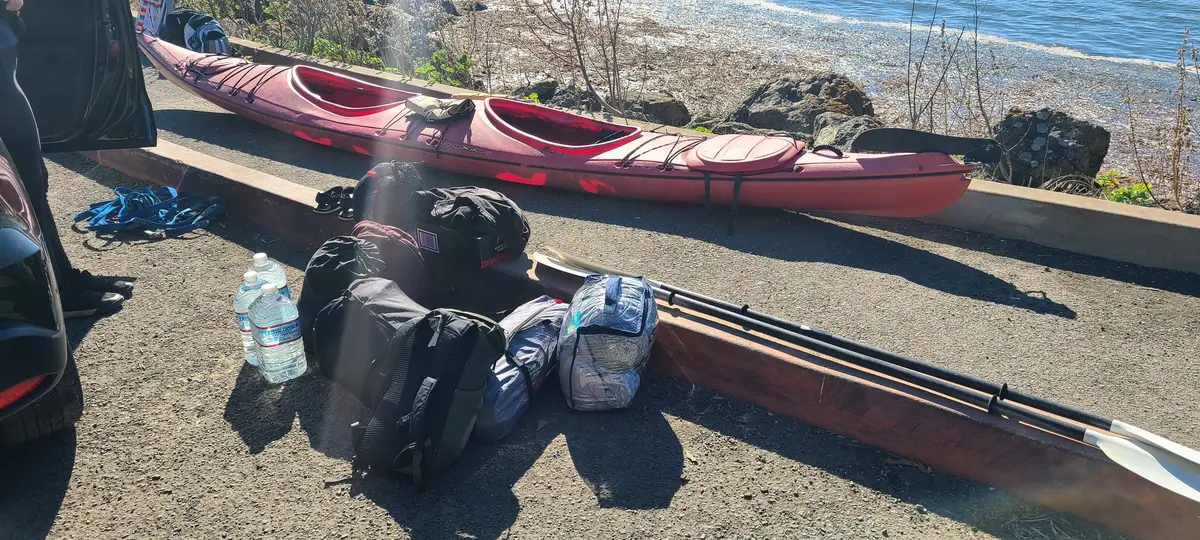
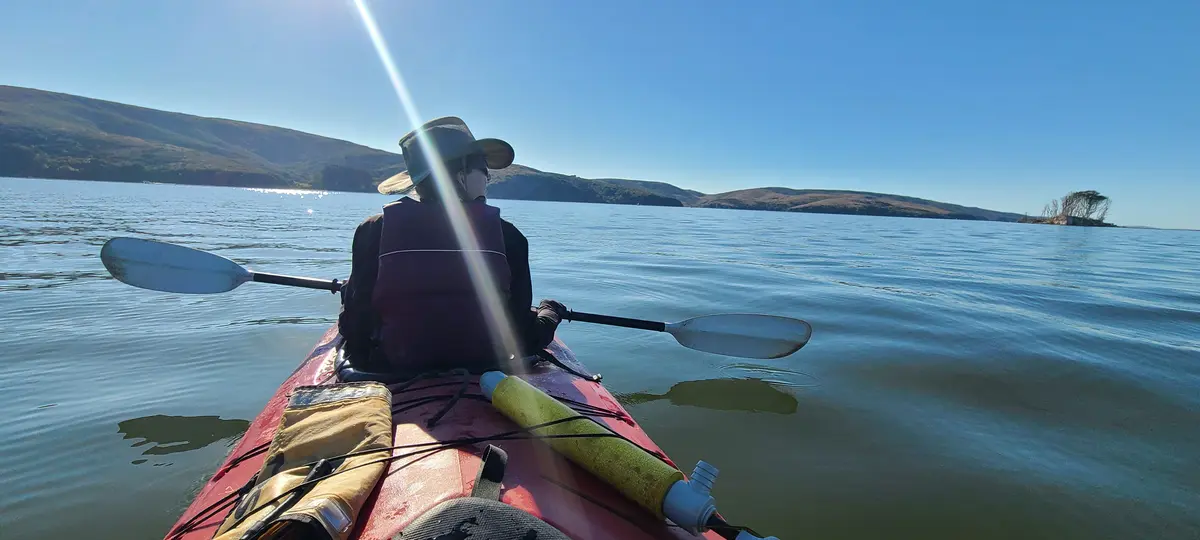
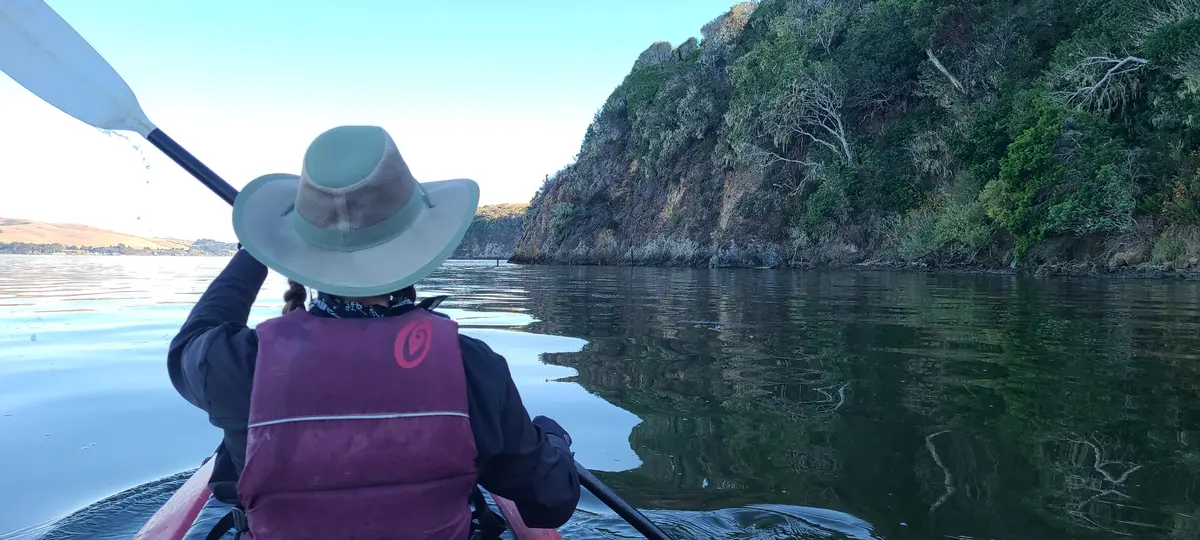
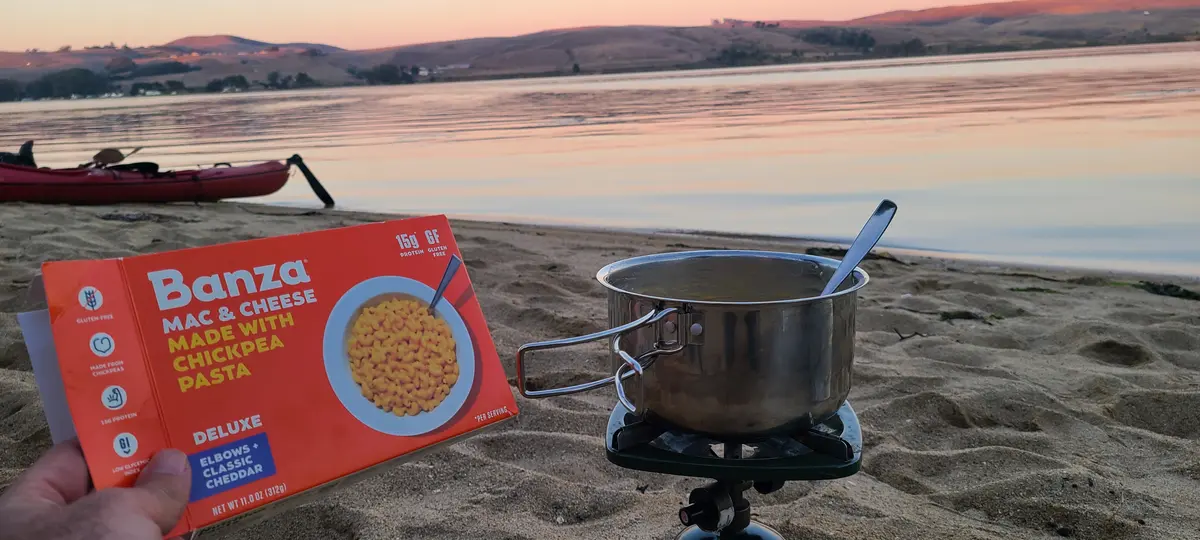




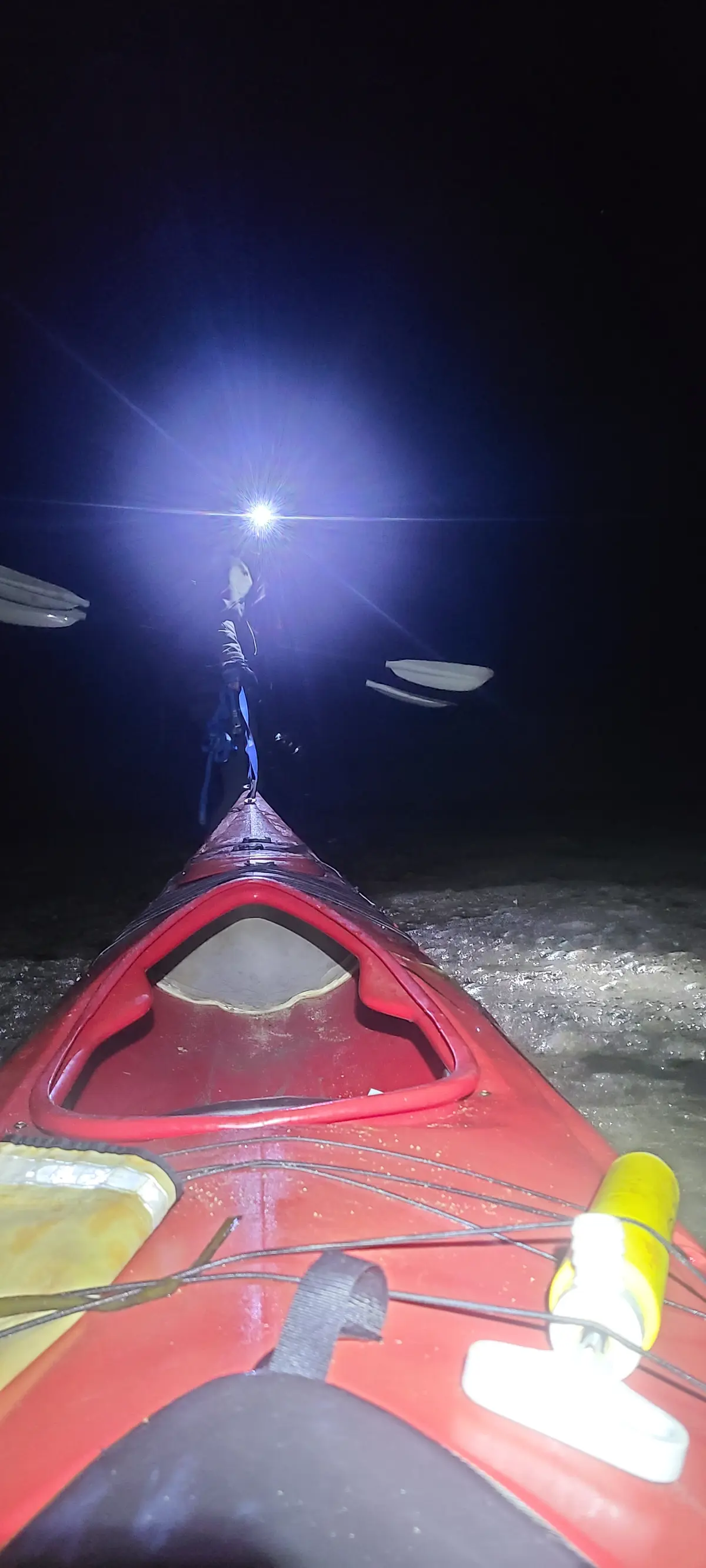
After dark, we got back on the water and kayaked from Tomales Beach up to White Gulch Beach - which is in a deep bay directly west of Hog Island. We found bioluminescence the whole way, and it was incredible. Every paddle stroke lit up the waters and disturbed kelp shot lightning bolts away from us.

Caveat to this post. There are several different ways to be dairy sensitive and lactose intolerance is only one of them. If you're eating dairy products that are supposed to be lactose-free and still having reactions a parsimonious explanation is that the lactose is not actually the problem. I'll discuss this in a future post.
Background
Lactose tolerance is common globally. Roughly two-thirds of the world's adults stop producing lactase past childhood, and what counts as a tolerable dose varies between people. The NIH puts the typical daily threshold at about 12 g of lactose — roughly one cup of milk — without symptoms or with only mild ones1. Below that, most people are fine; above it, dose and individual sensitivity start to matter.
The lactose content of dairy products spans nearly five orders of magnitude. A wedge of aged Cheddar has almost no lactose while a spoonful of Brunost (Norwegian whey cheese) has more than half its mass in lactose. The reason: lactose is water-soluble and lives in the whey, not the curd. Anything that drains whey (cheesemaking, Greek-style yogurt straining) removes lactose and anything that ferments whey (aging, live yogurt cultures) converts it to lactic acid. Anything that concentrates whey (Brunost, ricotta, dry milk solids) also concentrates lactose.
The tables
The table below is my personal reference. I've normalized all values to grams of lactose per 100 g of product, so they're directly comparable. Each value is footnoted. Where US and non-US values differ I've tried to go with the US value since I spend most of my time in the US.
For mental conversion: 100 g is about 1 cup of milk, about 3.5 oz of cheese, or roughly 7 tablespoons of butter.
The Safe? column estimates whether a lactose-intolerant adult can comfortably consume a typical US serving (1 cup milk/yogurt, 1 oz cheese, 1 tbsp butter, 2 tbsp cream, ½ cup ice cream, ½ cup cottage cheese / ricotta) with a 3× margin of safety. Since most people, myself included, eat more than one serving at a meal, a food is really only safe if ~3 servings still fit under the ~12 g/day NIDDK threshold1:
- Yes — < 1 g per typical serving. Three servings are still well under the 12g threshold. This is safe for nearly all lactose-intolerant adults.
- Maybe — 1–4 g per typical serving. One serving is fine for most people; 3 servings approach or hit the threshold.
- No — > 4 g per typical serving. Three servings clearly exceed the threshold; even one serving may cause symptoms in sensitive individuals.
Milks
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Whole cow milk (3.25% fat) | 4.812 | No | ≈ 12 g per 244 g cup1 — one cup alone hits threshold |
| Reduced-fat (2%) cow milk | 4.692 | No | |
| Lowfat (1%) cow milk | 4.862 | No | |
| Skim cow milk | 5.052 | No | Removing fat slightly raises lactose-by-mass |
| Buttermilk, lowfat (cultured) | 4.03 | No | Despite the name, US buttermilk is fermented lowfat milk |
| Goat milk | 4.272 | No | |
| Sheep milk | 4.44 | No | |
| Lactose-free milk (Lactaid-style) | <0.14 | Yes | Lactase enzyme has hydrolyzed the lactose into glucose + galactose |
| Evaporated whole milk | ~105 | Maybe | Used in small amounts (2 tbsp ≈ 3 g); a ½ cup recipe portion is "No" |
| Sweetened condensed milk | ~125 | Maybe | A 2 tbsp drizzle is fine; ¼ cup in a recipe is not |
Yogurts and fermented milks
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Plain whole-milk yogurt | 4.73 | No | A 6 oz container has ~8 g; live cultures pre-digest some lactose but most remains |
| Plain low-fat yogurt | 4.03 | No | ~6.8 g per 6 oz container |
| Greek yogurt (strained, plain) | ~3.06 | Maybe | Straining removes ~70% of the lactose into the acid whey6 |
| Greek yogurt (nonfat, strained) | ≤0.76 | Yes | |
| Kefir, plain | 4.03 | Maybe | High raw lactose, but live yeast/bacteria reduce symptoms in lactose-intolerant people by 54–71%7 — effective dose is much lower than the raw number suggests |
| Skyr | 2.53 | Maybe | Strained, like Greek yogurt |
| Sour cream | 2.03 | Yes | A 2 tbsp dollop has ~0.6 g |
| Crème fraîche | ~2.44 | Yes | A 2 tbsp serving has ~0.7 g |
Cream and butter
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Butter, salted | 0.68 | Yes | An entire stick (113 g) has ~0.1 g9; almost all the whey has been churned out |
| Butter oil | 0.0038 | Yes | |
| Ghee | 0.00298 | Yes | |
| Heavy / whipping cream | 2.53 | Yes | A 2 tbsp serving has ~0.75 g; more fat, less aqueous phase, less lactose |
| Half-and-half | ~4.05 | Maybe | A 2 tbsp coffee splash is fine (~1.2 g); 3 splashes approach the threshold |
Fresh and unripened cheeses
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Cottage cheese, low-fat | 1.83 | Maybe | A ½ cup serving has ~2 g; 3 servings = 6 g |
| Cream cheese | 3.63 | Maybe | Highest of any common cheese; a 1 oz schmear has ~1 g |
| Ricotta (whey-based) | 2.753 | Maybe | Made from the whey drained off other cheeses; ½ cup has ~3.4 g |
| Mascarpone | 3.03 | Yes | A 1 oz serving has ~0.84 g |
| Chèvre, fresh (goat) | 0.93 | Yes | A 1 oz serving has ~0.25 g |
| Paneer | 0.0023 | Yes | Acid-set; the whey carries off nearly all of it |
| Burrata | ~1.53 | Maybe | A mozzarella shell wrapped around a stracciatella cream filling; lactose is dominated by the filling. ~1 g per typical 1 oz serving |
| Stracciatella (burrata cream filling) | 1.83 | Maybe | Sold as a stand-alone topping; eaten by the spoonful |
| Cheese curds, fresh | 3.03 | Maybe | Wisconsin specialty. Unaged and unpressed, so much of the lactose remains; a small handful (~30 g) has ~1 g |
| Queso fresco | 2.310 | Maybe | Fresh, unaged Mexican cheese. Crumbled on tacos at typical portions (~30 g) gives <1 g |
Soft-ripened and washed-rind cheeses
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Brie | <0.00243 | Yes | Mould ripening fully ferments residual lactose |
| Camembert | <0.00243 | Yes | |
| Limburger | <0.00243 | Yes | |
| Taleggio | <0.0013 | Yes |
Pasta filata (stretched-curd) cheeses
| Product | Lactose (g/100g) | Safe? | Notes |
|---|---|---|---|
| Mozzarella, commercial low-moisture | 0.743 | Yes | The kind on US pizza |
| Mozzarella di bufala | 0.353 | Yes | |
| Bocconcini (mini fresh mozzarella) | <0.0013 | Yes | Fresh; lower lactose than commercial mozzarella because there's less moisture to retain whey |
| String cheese | 0.743 | Yes | Mechanically the same product as low-moisture mozzarella; one stick (~28 g) has ~0.2 g |
| Oaxaca (queso Oaxaca) | <0.53 | Yes | Mexican pasta filata cheese; USDA reports ~0 carbohydrates per serving |
| Provolone, dolce or piccante | <0.0013 | Yes |
My current and previous choices for the tools I use.
- Android Apps
- Workout tracker: FitBod (but probably not for much longer - it doesn't have a data export option)
- Diet tracker: MacroFactor
- Bathroom
- Toothbrush: Philips One by Sonicare - compact, lightweight, good battery, charges with a USB-A to USB-C cable. The only improvement here would be the ability to charge with a real USB-C cable.
- Fingernail clipper: Seki Edge
- Nail file: 3 Swords Germany
- Electronics
- Large power adapter: Anker Prime 67W USB C Charger
- Small power adapter: Anker Nano Charger
- Wireless charger: TOZO W1 Wireless Charger 15W Max
- USB-C cable: Anker
- Wireless car charger: APPS2Car
- Car USB-C charger: Roadress
- USB-C magnetic breakaways: TiMOVO
- Keyboard: nuphy Air75
- USB-C to USB-A adapter: Syntech
- Computer camera cover: CloudValley
- Hiking
- Climbing
- Belay glasses: BG Climbing
- PPE
- Games (Best)
- Social Games
- Games (interesting)
- Misc
- Micro-knife - has passed through TSA many times unmolested
- Kitchen
An answer to this question on Stack Overflow.
Question
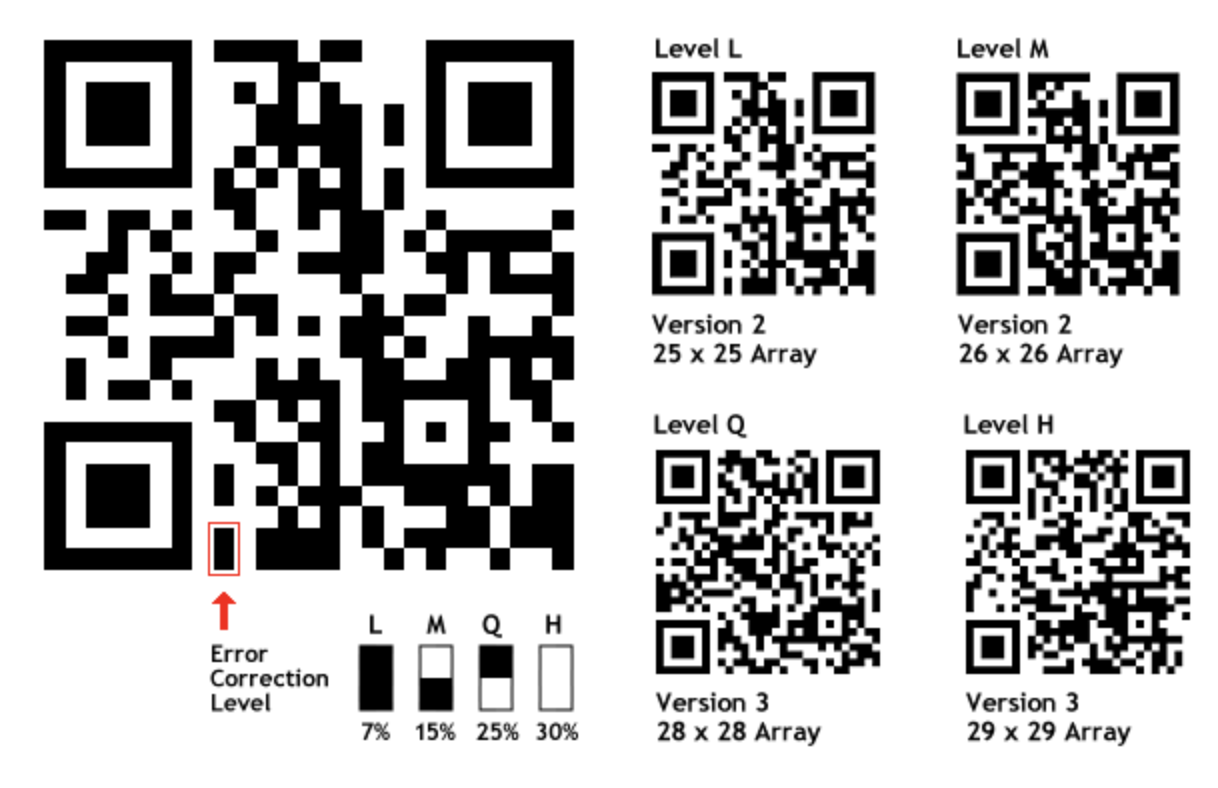
I'm not a specialist, but as far as I know, a bit of information in a QR-code is coded more than once, and it is defined as the redundancy level
How can I estimate a QR-code redundancy level ? Is where an mobile app or a website where I can test my QR-code redundancy level easily ? If not, is it an easy algorithm that I can implement ?
Redundancy is sorted in different categories according to this website, but I'd like to have the direct percentage value if possible
Answer
QR codes contain a couple of bits which indicate the error correction level, as depicted below (source):
An answer to this question on Stack Overflow.
Question
The sigmoid function is defined as
S(t) = 1 / (1 + e^(-t))
(where ^ is pow)
I found that using the C built-in function exp() to calculate the value of f(x) is slow. Is there any faster algorithm to calculate the value of f(x)?
Answer
This question has significantly more detail including these benchmarked results:
name rms_error maxdiff time_us speedup samples
logistic_with_tanh 5.9496e-02 1.5014e-01 0.0393 0.5076 200000001
logistic_with_atan 3.9051e-02 9.6934e-02 0.0321 0.6211 200000001
logistic_with_erf 6.5068e-02 1.6581e-01 0.0299 0.6676 200000001
logistic_fexp_quintic_approx 1.2921e-07 5.9050e-07 0.0246 0.8118 200000001
logistic_product_approx_float128 8.7032e-04 1.7217e-03 0.0209 0.9523 200000001
logistic_with_exp_no_overflow 4.7660e-17 1.6653e-16 0.0198 1.0084 200000001
logistic_product_approx128 8.7032e-04 1.7211e-03 0.0164 1.2187 200000001
log_w_approx_exp_no_overflow128 8.7193e-04 1.7211e-03 0.0158 1.2640 200000001
logistic_with_sqrt 8.3414e-02 1.1086e-01 0.0146 1.3662 200000001
log_w_approx_exp_no_overflow16 6.9726e-03 1.4074e-02 0.0141 1.4114 200000001
log_w_approx_exp_no_overflow16_clamped 6.9726e-03 1.4074e-02 0.0141 1.4153 200000001
logistic_schraudolph_approx 1.5661e-03 8.9906e-03 0.0138 1.4497 200000001
logistic_with_abs 6.0968e-02 8.2289e-02 0.0134 1.4936 200000001
logistic_orig 0.0000e+00 0.0000e+00 0.0199 ------ 200000001