Maximum volume inscribed ellipsoid in a polytope/set of points
An answer to this question on Stack Overflow.
Question
Later Edit: I uploaded here a sample of my original data. It's actually a segmentation image in the DICOM format. The volume of this structure as it is it's ~ 16 mL, so I assume the inner ellipsoid volume should be smaller than that. to extract the points from the DICOM image I used the following code:
import os
import numpy as np
import SimpleITK as sitk
def get_volume_ml(image):
x_spacing, y_spacing, z_spacing = image.GetSpacing()
image_nda = sitk.GetArrayFromImage(image)
imageSegm_nda_NonZero = image_nda.nonzero()
num_voxels = len(list(zip(imageSegm_nda_NonZero[0],
imageSegm_nda_NonZero[1],
imageSegm_nda_NonZero[2])))
if 0 >= num_voxels:
print('The mask image does not seem to contain an object.')
return None
volume_object_ml = (num_voxels * x_spacing * y_spacing * z_spacing) / 1000
return volume_object_ml
def get_surface_points(folder_path):
"""
:param folder_path: path to folder where DICOM images are stored
:return: surface points of the DICOM object
"""
# DICOM Series
reader = sitk.ImageSeriesReader()
dicom_names = reader.GetGDCMSeriesFileNames(os.path.normpath(folder_path))
reader.SetFileNames(dicom_names)
reader.MetaDataDictionaryArrayUpdateOn()
reader.LoadPrivateTagsOn()
try:
dcm_img = reader.Execute()
except Exception:
print('Non-readable DICOM Data: ', folder_path)
return None
volume_obj = get_volume_ml(dcm_img)
print('The volume of the object in mL:', volume_obj)
contour = sitk.LabelContour(dcm_img, fullyConnected=False)
contours = sitk.GetArrayFromImage(contour)
vertices_locations = contours.nonzero()
vertices_unravel = list(zip(vertices_locations[0], vertices_locations[1], vertices_locations[2]))
vertices_list = [list(vertices_unravel[i]) for i in range(0, len(vertices_unravel))]
surface_points = np.array(vertices_list)
return surface_points
folder_path = r"C:\Users\etc\TTT [13]\20160415 114441\Series 052 [CT - Abdomen WT 1 0 I31f 3]"
points = get_surface_points(folder_path)
I have a set of points (n > 1000) in 3D space that describe a hollow ovoid like shape. What I would like is to fit an ellipsoid (3D) that is inside all of the points. I am looking for the maximum volume ellipsoid fitting inside the points.
I tried to adapt the code from Minimum Enclosing Ellipsoid (aka outer bounding ellipsoid)
by modifying the threshold err > tol, with my logic begin that all points should be smaller than < 1 given the ellipsoid equation. But no success.
I also tried the Loewner-John adaptation on mosek, but I didn't figure how to describe the intersection of a hyperplane with 3D polytope (the Ax <= b representation) so I can use it for the 3D case. So no success again.
The code from the outer ellipsoid:
import numpy as np
import numpy.linalg as la
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
pi = np.pi
sin = np.sin
cos = np.cos
def plot_ellipsoid(A, centroid, color, ax):
"""
:param A: matrix
:param centroid: center
:param color: color
:param ax: axis
:return:
"""
centroid = np.asarray(centroid)
A = np.asarray(A)
U, D, V = la.svd(A)
rx, ry, rz = 1. / np.sqrt(D)
u, v = np.mgrid[0:2 * np.pi:20j, -np.pi / 2:np.pi / 2:10j]
x = rx * np.cos(u) * np.cos(v)
y = ry * np.sin(u) * np.cos(v)
z = rz * np.sin(v)
E = np.dstack((x, y, z))
E = np.dot(E, V) + centroid
x, y, z = np.rollaxis(E, axis=-1)
ax.plot_wireframe(x, y, z, cstride=1, rstride=1, color=color, alpha=0.2)
ax.set_zlabel('Z-Axis')
ax.set_ylabel('Y-Axis')
ax.set_xlabel('X-Axis')
def mvee(points, tol = 0.001):
"""
Finds the ellipse equation in "center form"
(x-c).T * A * (x-c) = 1
"""
N, d = points.shape
Q = np.column_stack((points, np.ones(N))).T
err = tol+1.0
u = np.ones(N)/N
while err > tol:
# assert u.sum() == 1 # invariant
X = np.dot(np.dot(Q, np.diag(u)), Q.T)
M = np.diag(np.dot(np.dot(Q.T, la.inv(X)), Q))
jdx = np.argmax(M)
step_size = (M[jdx]-d-1.0)/((d+1)*(M[jdx]-1.0))
new_u = (1-step_size)*u
new_u[jdx] += step_size
err = la.norm(new_u-u)
u = new_u
c = np.dot(u,points)
A = la.inv(np.dot(np.dot(points.T, np.diag(u)), points)
- np.multiply.outer(c,c))/d
return A, c
folder_path = r"" # path to a DICOM img folder
points = get_surface_points(folder_path) # or some random pts
A, centroid = mvee(points)
U, D, V = la.svd(A)
rx_outer, ry_outer, rz_outer = 1./np.sqrt(D)
# PLOT
fig = plt.figure()
ax1 = fig.add_subplot(111, projection='3d')
ax1.scatter(points[:, 0], points[:, 1], points[:, 2], c='blue')
plot_ellipsoid(A, centroid, 'green', ax1)
Which gives me this result for the outer ellipsoid on my sample points:
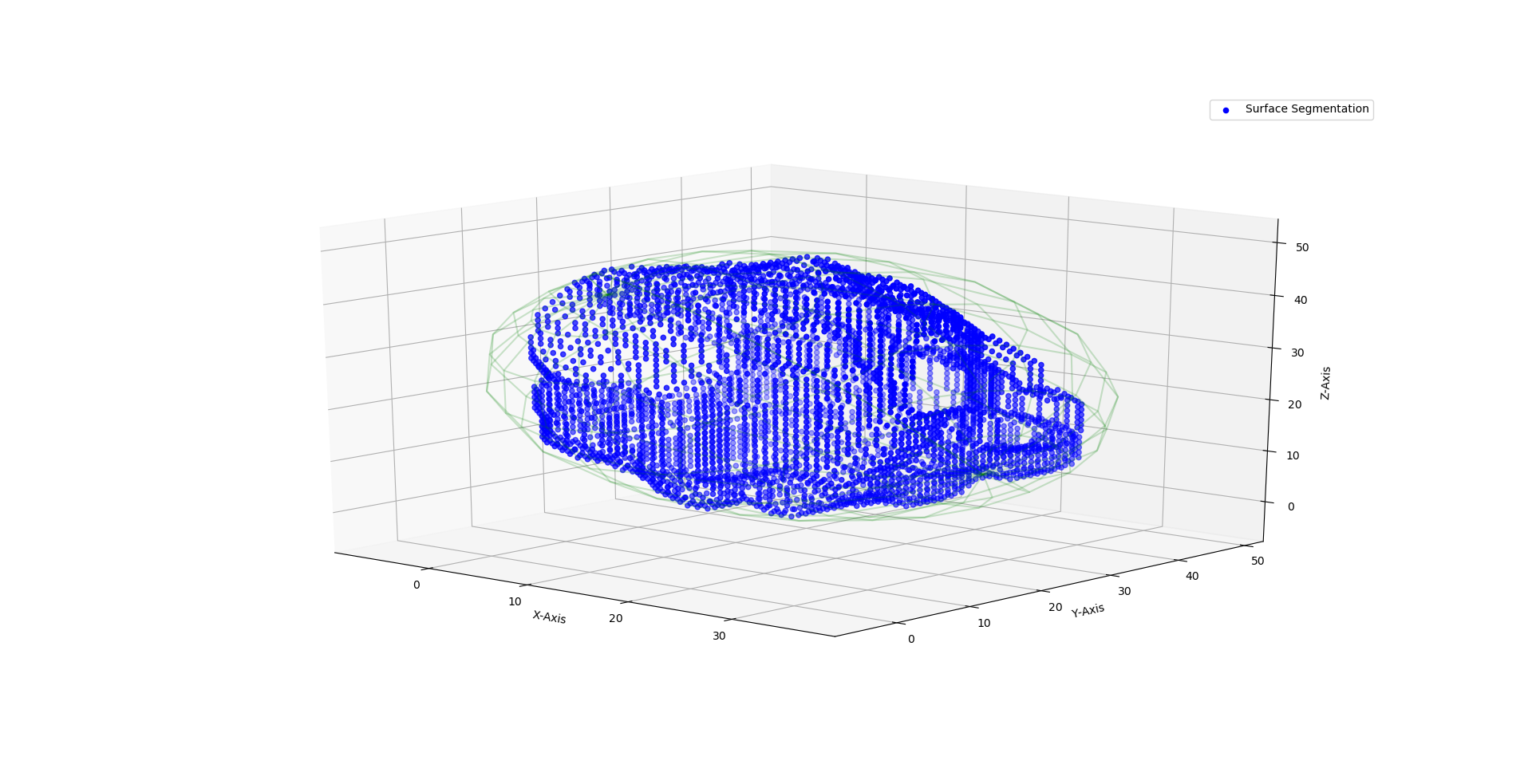

The main question: How do I fit an ellipsoid (3D) inside a cloud of 3D points using Python?
Is it possible to modify the algorithm for the outer ellipsoid to get the maximum inscribed (inner) ellipsoid?
I am looking for code in Python ideally.
Answer
Whether this answer works depends on how much noise is in your data. The idea is to first find the point cloud's convex hull and then find the largest ellipsoid which fits within this hull. If most of your points lie close to the surface of the ellipsoid they describe than this approximation won't be "too bad".
To do so, note that a convex hull can be described by a set of linear inequalities Ax<=b.
Note that the bounding ellipsoid can be described by E={Bx+d for ||x||_2<=1}, where B is a positive semi-definite matrix describing how and which directions the ellipsoid is stretched and d is a vector describing its offset.
Note that the volume of the ellipsoid is given by det(B^-1). If we were to try to maximize or minimize this determinant we would fail because that would give a non-convex problem. However, applying a log transform log(det(B^-1)) makes the problem convex again. The optimization program we're going to use doesn't allow matrix inverses, but it's easy to show that the foregoing is equivalent to -log(det(B)).
Finally, some bracing algebraic manipulation gives us the optimization problem:
minimize -log(det(B))
s.t. ||B*A_i||_2 + a_i^T * d <= b_i, i = 1, ..., m
B is PSD
We can solve this in Python using CVXPY as follows:
#!/usr/bin/env python3
from mpl_toolkits.mplot3d import axes3d
from scipy.spatial import ConvexHull
import cvxpy as cp
import matplotlib.pyplot as plt
import numpy as np
import sklearn.datasets
#From: https://stackoverflow.com/a/61786434/752843
def random_point_ellipsoid(a,b,c,x0,y0,z0):
"""Generate a random point on an ellipsoid defined by a,b,c"""
u = np.random.rand()
v = np.random.rand()
theta = u * 2.0 * np.pi
phi = np.arccos(2.0 * v - 1.0)
sinTheta = np.sin(theta);
cosTheta = np.cos(theta);
sinPhi = np.sin(phi);
cosPhi = np.cos(phi);
rx = a * sinPhi * cosTheta;
ry = b * sinPhi * sinTheta;
rz = c * cosPhi;
return rx, ry, rz
def random_point_ellipse(W,d):
# random angle
alpha = 2 * np.pi * np.random.random()
# vector on that angle
pt = np.array([np.cos(alpha),np.sin(alpha)])
# Ellipsoidize it
return W@pt+d
def GetRandom(dims, Npts):
if dims==2:
W = sklearn.datasets.make_spd_matrix(2)
d = np.array([2,3])
points = np.array([random_point_ellipse(W,d) for i in range(Npts)])
elif dims==3:
points = np.array([random_point_ellipsoid(3,5,7,2,3,3) for i in range(Npts)])
else:
raise Exception("dims must be 2 or 3!")
noise = np.random.multivariate_normal(mean=[0]*dims, cov=0.2*np.eye(dims), size=Npts)
return points+noise
def GetHull(points):
dim = points.shape[1]
hull = ConvexHull(points)
A = hull.equations[:,0:dim]
b = hull.equations[:,dim]
return A, -b, hull #Negative moves b to the RHS of the inequality
def Plot(points, hull, B, d):
fig = plt.figure()
if points.shape[1]==2:
ax = fig.add_subplot(111)
ax.scatter(points[:,0], points[:,1])
for simplex in hull.simplices:
plt.plot(points[simplex, 0], points[simplex, 1], 'k-')
display_points = np.array([random_point_ellipse([[1,0],[0,1]],[0,0]) for i in range(100)])
display_points = display_points@B+d
ax.scatter(display_points[:,0], display_points[:,1])
elif points.shape[1]==3:
ax = fig.add_subplot(111, projection='3d')
ax.scatter(points[:,0], points[:,1], points[:,2])
display_points = np.array([random_point_ellipsoid(1,1,1,0,0,0) for i in range(1000)])
display_points = display_points@B+d
ax.scatter(display_points[:,0], display_points[:,1], display_points[:,2])
plt.show()
def FindMaximumVolumeInscribedEllipsoid(points):
"""Find the inscribed ellipsoid of maximum volume. Return its matrix-offset form."""
dim = points.shape[1]
A,b,hull = GetHull(points)
B = cp.Variable((dim,dim), PSD=True) #Ellipsoid
d = cp.Variable(dim) #Center
constraints = [cp.norm(B@A[i],2)+A[i]@d<=b[i] for i in range(len(A))]
prob = cp.Problem(cp.Minimize(-cp.log_det(B)), constraints)
optval = prob.solve()
if optval==np.inf:
raise Exception("No solution possible!")
print(f"Optimal value: {optval}")
Plot(points, hull, B.value, d.value)
return B.value, d.value
FindMaximumVolumeInscribedEllipsoid(GetRandom(dims=2, Npts=100))
FindMaximumVolumeInscribedEllipsoid(GetRandom(dims=3, Npts=100))
Solutions are computed quickly.
Visually, this gives (for 2D):
Note that I've added a lot of noise to emphasize what's going on.
And for 3D:
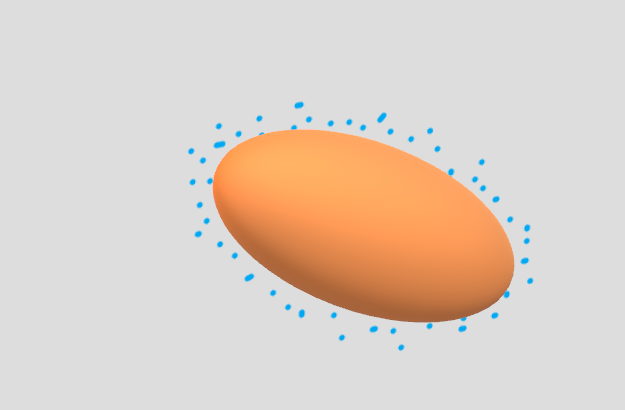
Although the code above is written for two or three dimensions, you could easily adapt it for any number of dimensions, though the visualization will become more difficult.
If the convex hull is not good and you want some kind of "interior convex hull", that'll be harder: this hull isn't well-defined. However, you could use alpha shapes to try to find such a hull and then use the algorithm above to solve for it.
Note also that since we're using a convex polytope to bound the ellipse, rather than the points themselves, even if the points perfectly described an ellipsoid, we end up with an underestimated volume. We can visualize this, as below:
If the vertices of the square are the points, then the square is their convex hull. The circle bounded by the hull is clearly smaller than the circle that would be bounded only by the points.
EDIT: TO get the volume, you need to convert pixel indices to the coordinate system of your DICOM image, like so (NOTE: I'm not sure if I've scaled the correct coordinates by the correct values, but you'll be able to figure this out given your knowledge of the data):
from mpl_toolkits.mplot3d import axes3d
from scipy.spatial import ConvexHull
import cvxpy as cp
import matplotlib.pyplot as plt
import numpy as np
import os
import sklearn.datasets
import SimpleITK as sitk
import code
def get_volume_ml(image):
x_spacing, y_spacing, z_spacing = image.GetSpacing()
image_nda = sitk.GetArrayFromImage(image)
imageSegm_nda_NonZero = image_nda.nonzero()
num_voxels = len(list(zip(imageSegm_nda_NonZero[0],
imageSegm_nda_NonZero[1],
imageSegm_nda_NonZero[2])))
if 0 >= num_voxels:
print('The mask image does not seem to contain an object.')
return None
volume_object_ml = (num_voxels * x_spacing * y_spacing * z_spacing) / 1000
return volume_object_ml
def get_surface_points(dcm_img):
x_spacing, y_spacing, z_spacing = dcm_img.GetSpacing()
contour = sitk.LabelContour(dcm_img, fullyConnected=False)
contours = sitk.GetArrayFromImage(contour)
vertices_locations = contours.nonzero()
vertices_unravel = list(zip(vertices_locations[0], vertices_locations[1], vertices_locations[2]))
vertices_list = [list(vertices_unravel[i]) for i in range(0, len(vertices_unravel))]
surface_points = np.array(vertices_list)
surface_points = surface_points.astype(np.float64)
surface_points[:,0] *= x_spacing/10
surface_points[:,1] *= y_spacing/10
surface_points[:,2] *= z_spacing/10
return surface_points
def get_dcm_image(folder_path):
reader = sitk.ImageSeriesReader()
dicom_names = reader.GetGDCMSeriesFileNames(os.path.normpath(folder_path))
reader.SetFileNames(dicom_names)
reader.MetaDataDictionaryArrayUpdateOn()
reader.LoadPrivateTagsOn()
try:
dcm_img = reader.Execute()
except Exception:
raise Exception('Non-readable DICOM Data: ', folder_path)
return dcm_img
def GetHull(points):
dim = points.shape[1]
hull = ConvexHull(points)
A = hull.equations[:,0:dim]
b = hull.equations[:,dim]
return A, -b, hull #Negative moves b to the RHS of the inequality
def FindMaximumVolumeInscribedEllipsoid(points):
"""Find the inscribed ellipsoid of maximum volume. Return its matrix-offset form."""
dim = points.shape[1]
A,b,hull = GetHull(points)
B = cp.Variable((dim,dim), PSD=True) #Ellipsoid
d = cp.Variable(dim) #Center
constraints = [cp.norm(B@A[i],2)+A[i]@d<=b[i] for i in range(len(A))]
prob = cp.Problem(cp.Minimize(-cp.log_det(B)), constraints)
optval = prob.solve()
if optval==np.inf:
raise Exception("No solution possible!")
print(f"Optimal value: {optval}")
return B.value, d.value
#From: https://stackoverflow.com/a/61786434/752843
def random_point_ellipsoid(a,b,c,x0,y0,z0):
"""Generate a random point on an ellipsoid defined by a,b,c"""
u = np.random.rand()
v = np.random.rand()
theta = u * 2.0 * np.pi
phi = np.arccos(2.0 * v - 1.0)
sinTheta = np.sin(theta);
cosTheta = np.cos(theta);
sinPhi = np.sin(phi);
cosPhi = np.cos(phi);
rx = a * sinPhi * cosTheta;
ry = b * sinPhi * sinTheta;
rz = c * cosPhi;
return rx, ry, rz
def Plot(points, B, d):
hull = ConvexHull(points)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.scatter(points[:,0], points[:,1], points[:,2], marker=".")
display_points = np.array([random_point_ellipsoid(1,1,1,0,0,0) for i in range(1000)])
display_points = display_points@B+d
ax.scatter(display_points[:,0], display_points[:,1], display_points[:,2])
plt.show()
folder_path = r"data"
dcm_img = get_dcm_image(folder_path)
points = get_surface_points(dcm_img)
B, d = FindMaximumVolumeInscribedEllipsoid(points)
Plot(points, B, d)
ball_vol = 4/3.0*np.pi*(1.0**3)
print("DCM vol: ", get_volume_ml(dcm_img))
print("Ellipsoid Volume: ", np.linalg.det(B) * ball_vol)
This gives
DCM vol: 16.2786318359375
Ellipsoid Volume: 11.947614772444393