How do I align peak labels?
An answer to this question on Stack Overflow.
Question
I am attempting to graph data with peaks in R. The graphing itself has progressed well, but I've run into issues with labelling the relevant peaks. My current labelling system, detailed below, shifts the peak labels oddly to the side and results in lines crossing each other. Is there a way to align labels with the peaks themselves, or otherwise organize them aesthetically?
The following code reproduces my problem, using this data.
library(ggplot2)
library(ggpmisc)
library(ggrepel)
x=read.csv("data.csv")
colnames(x)=c("wv", "abs")
ggplot(x, aes(x=wv, y=abs)) + geom_line() + xlab(bquote('Wavenumbers ('~cm^-1*')')) + ylab("Absorbance (A.U.)") + scale_x_reverse(limits=c(2275,1975), expand=c(0,0)) + ylim(-0.01,0.29) + stat_peaks(colour = "black", span = 11, geom ="text_repel", direction = "y", angle = 90, ignore_threshold = 0.09, size = 3, x.label.fmt = "%.2f", vjust = 1, hjust = 0, segment.color = "red") + ggtitle("FTIR - Carbon Monoxide, Fundamentals")
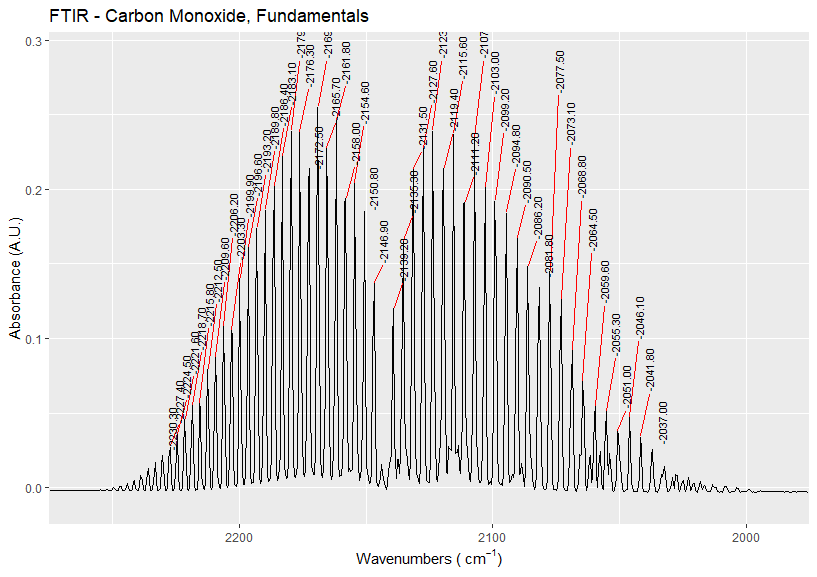
Answer
I wasn't able to find a way to do this without constructing an auxiliary dataset. However, once constructed the actual plotting then becomes fairly easy:
The code is as follows:
library(dplyr)
library(ggplot2)
library(ggpmisc)
library(ggrepel)
library(pracma)
#####################################
# Constants
#####################################
wvfilter = c(1975, 2275)
data_filename = "data.csv"
#####################################
# Read data
#####################################
dat = read.csv(data_filename)
colnames(dat)=c("wv", "abs")
#####################################
# Identify peaks
#####################################
# Extract peaks
peaks = data.frame(findpeaks(dat$abs, threshold=0, minpeakheight=0.03))
# Give data frame reasonable names
colnames(peaks) = c("height", "x_index", "peak_begin_index", "peak_end_index")
# Convert from index to wavelength
peaks$wv = dat$wv[peaks$x_index]
# Set the y position of the labels
peaks$nudge = 0.3-peaks$height
# Filter by wavelength
peaks = peaks %>% filter(wvfilter[1] <= wv & wv <= wvfilter[2])
#####################################
# Plot data
#####################################
ggplot(x, aes(x=wv, y=abs)) +
geom_line() +
geom_text_repel(
data=peaks,
mapping=aes(x=wv, y=height, label=height),
force=0,
nudge_y=peaks$nudge,
direction="x",
angle=90,
segment.color="red"
) +
xlab(bquote('Wavenumbers ('~cm^-1*')')) +
ylab("Absorbance (A.U.)") +
scale_x_reverse(limits=rev(wvfilter), expand=c(0,0)) +
ylim(-0.01,0.29) +
ggtitle("FTIR - Carbon Monoxide, Fundamentals")
Some notes:
In the image I'm using the wrong text for the labels. Fix this with
label=wvinstead oflabel=height.The
peaksdataset needs to be filtered using the same values passed toscale_x_reverse(limits=. To simplify this, I've introduced a global variable for setting the x-axis limits.Playing with the
forceargument gives different degrees of clustering of the labels.force=0seems to work best for your data.You need to manually set the
peaks$nudgevalue. I don't know of a good way of doing this other than guess-and-check.