Approximating the boundary between two sets of points (in 2D): Fitting a region
An answer to this question on the Scientific Computing Stack Exchange.
Question
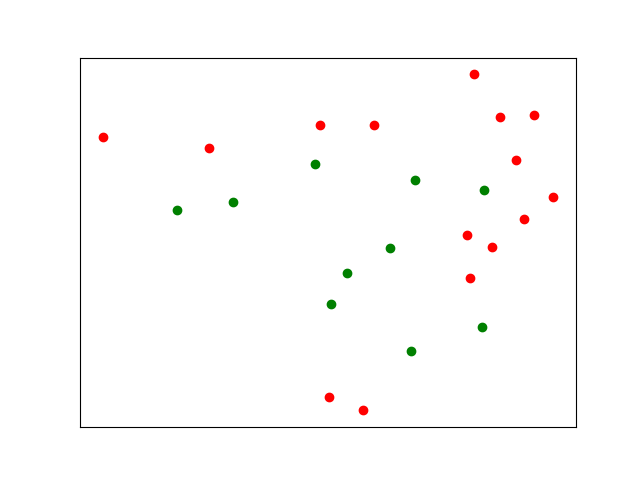
Given two sets of points $p_{\text{in},i}$ and $p_{\text{out},j}$ inside and outside of what I intuitively call a "region", I would like to estimate and describe the boundary of this region. Two kinds of answer are welcome:
- A parametrized description of the boundary
- A estimation function that predicts whether a new point is inside the region.
So far, I have found lots of clustering algorithms, which may be related, but I do not see how to apply them here. Here, in/out-ness is part of the input, not the result.
I arrived at this question from an experimental context. A nice follow-up question is "where to probe next" (i.e. at which new location would in/out-knowledge change the model most? In the image below: bottom left)
My first naive approach is something like "a circle that minimizes some kind of signed distance to all points", but I want more complex shapes.
Some more thoughts:
- I'm solely interested in 2D right now
- I'm fairly proficient at optimization, even pointers to suitable parametrizable "descriptions of regions" would be welcome. I know these two sequences, for example, but they don't seem suitably bounded.
- The number of optimization parameters should be scalable: for just one point in and one point inside, even returning a circle is overfitting, but for the data below, a more complex shape of the region makes sense.
- There will be outliers
- An alogrithm for convex regions only would be acceptable, but a more general one is preferred.
- Weights (uncertainties) for the points are available.
Updates
My original question was unclear in some ways. Thanks to the first answers, I can now specify some things more clearly:
- "What I intuitively call a region" has two important properties: it is finite and connected
- There will be outliers: Some points will be wrongly classified, and the model shall not overfit to include all points. Hence "number of parameters should be scalable" to avoid/adjust overfitting, just like one will usually not fit a polynomial of degree $n$ to $n-1$ points.
Answer
Theory
Clustering is unlikely to work in this case because your red points are separated from each other by the green points. You could use more clusters, but this will require a lot of manual inspection and fiddling.
A standard approach to this sort of problem is to use a nonlinear support vector machine. The idea, simply described, is that although your data may not be separable in two dimensions projecting the data into a higher dimension always guarantees a clean separator which can then be used to classify future points of interest. Furthermore, we can choose this separator so that the distance between the separator and any of the data points is maximized.
Here are a couple of images showing the idea:
Results
I used WebPlotDigitizer to extract your data in arbitrary, but aspect-ratio-preserving units and then ran this through a support-vector machine using a radial basis function kernel. This gives the following separator:
Notice that, as promised, this cleanly separates your data into two classes. The separator itself is the solid grey line while the margins around the separator (the distance between the separator and the data) are shown with dotted lines. The distance between the separator and these margins is the maximum versus all other choices for this kernel. The data points which induce the margins are circled.
If we add a hundred new points and classify them using our SVM, we get this:
Implementation
I use Python to implement the above ideas.
import matplotlib.pyplot as plt
import numpy as np
from sklearn import svm
# Your data
data = [
[ 0.7490599550363091 , 9.24443743612264 , 0],
[ 4.12034765121126 , 8.885539916739752 , 0],
[ 7.6539862433328665 , 9.612037680925432 , 0],
[ 7.914094745271155 , 0.9253511681985884 , 0],
[ 8.994538029638242 , 0.5077641947442704 , 0],
[ 9.371974940345583 , 9.608153819994236 , 0],
[12.320128383367402 , 6.088728506174032 , 0],
[12.411426660363531 , 4.715352126892222 , 0],
[12.558352293037347 , 11.229593835418646 , 0],
[13.114336626127825 , 5.703722810531428 , 0],
[13.381386496538893 , 9.854563219073547 , 0],
[13.886274645038084 , 8.480251836234228 , 0],
[14.13510342320811 , 6.595572357695316 , 0],
[14.463275899337807 , 9.915985760466931 , 0],
[15.05984795641495 , 7.296033868971857 , 0],
[ 3.096302985685499 , 6.907935469254409 , 1],
[ 4.878692156862327 , 7.159379502874164 , 1],
[ 7.4911533079089345 , 8.366972559074298 , 1],
[ 7.986689890548966 , 3.89506631355318 , 1],
[ 8.498712223311847 , 4.883868537295852 , 1],
[ 9.869150457208352 , 5.679125024633843 , 1],
[10.527327159481406 , 2.3884159656508306 , 1],
[10.671071331605198 , 7.848836741512311 , 1],
[12.788477939489049 , 3.1497246315161327 , 1],
[12.86531503216689 , 7.524534353757313 , 1],
]
# Load data into numpy
data = np.array(data)
# Separate data into x and y values; predictors and observations
data_x = data[:,0:2] # x-value/predictor
data_y = data[:,2] # y-value/observation
# In these "learning" style problems you often want to avoid over-fitting, so
# it might make sense to split your data into training and test sets. Here we
# simply use all data as the training data.
x_train = data_x[:,:]
y_train = data_y[:]
# Using the RBF (radial basis function) kernel with an SVM projects the points
# into a higher-dimensional space where a clean boundary is possible and then
# lowers that boundary back into the space the points live in.
model = svm.SVC(kernel='rbf', C=1e6)
model.fit(x_train, y_train)
def plot_model(data_x, data_y, predict_x=None, predict_y=None) -> None:
# Let's plot some stuff
fig, ax = plt.subplots(figsize=(12, 7))
# Create grid to evaluate model
xx = np.linspace(-1, max(data_x[:,0]) + 1, len(data_x))
yy = np.linspace(0, max(data_x[:,1]) + 1, len(data_x))
YY, XX = np.meshgrid(yy, xx)
xy = np.vstack([XX.ravel(), YY.ravel()]).T
# Assigning different colors to the classes
colors = data_y
colors = np.where(colors == 1, '#8C7298', '#4786D1')
ax.scatter(data_x[:,0], data_x[:,1],c=colors)
if predict_x is not None:
assert predict_y is not None
colors = predict_y
colors = np.where(colors == 1, '#8C7298', '#4786D1')
ax.scatter(predict_x[:,0], predict_x[:,1],c=colors,s=4)
# Get the separator
Z = model.decision_function(xy).reshape(XX.shape)
# Draw the decision boundary and margins
ax.contour(XX, YY, Z, colors='k', levels=[-1, 0, 1], alpha=0.5, linestyles=['--', '-', '--'])
# Highlight support vectors with a circle around them
ax.scatter(model.support_vectors_[:, 0], model.support_vectors_[:, 1], s=100, linewidth=1, facecolors='none', edgecolors='k')
plot_model(data_x, data_y)
plt.show()
# Generate some previously-unseen data
new_x = np.random.uniform(low=min(data_x[:,0]), high=max(data_x[:,0]), size=(100,1))
new_y = np.random.uniform(low=min(data_x[:,1]), high=max(data_x[:,1]), size=(100,1))
new_data = np.hstack((new_x, new_y))
predictions_for_new_data = model.predict(new_data)
plot_model(data_x, data_y, new_data, predictions_for_new_data)
plt.show()